RECENT |
April 6, 2024
March 16, 2024
March 2, 2024

MOVIE NIGHT We watched the 2011 movie by Discovery Institute, "DARWIN'S HERETIC - Did the Co-Founder of Evolution Embrace Intelligent Design?"
From the DVD cover: Alfred Russel Wallace was one of the most renowned biologists of the nineteenth and early twentieth centuries. He shares credit with Charles Darwin for developing the theory of evolution by natural selection, and is widely regarded as the father of the field of biogeography. Dubbed by historian, Michael Flannery a "Victorian Indiana Jones," Wallace explored the Amazon, lived with headhunters, and even survived a shipwreck.
Yet one facet of Wallace's remarkable career has been ignored: His embrace of intelligent design.
The story of Wallace's doubts about the creative power of natural selection and his growing belief that nature requires an intelligent cause is one of the forgotten chapters in the history of science. But it's an intriguing story that sheds new light on current debates. In this DVD, take a fascinating journey with Wallace as he breaks with Darwin and comes to believe that nature is permeated with evidence of purposeful design.
(Full Runtime, including extra features, 54 minutes.)
The primary portion of this video, the 21-minute documentary section, streams live on YouTube, at https://www.youtube.com/watch?v=hxvAVln6HLI.
The DVD is available from Amazon for $6.61
This meeting WAS NOT be recorded or broadcast to a ZOOM audience.
From the DVD cover: Alfred Russel Wallace was one of the most renowned biologists of the nineteenth and early twentieth centuries. He shares credit with Charles Darwin for developing the theory of evolution by natural selection, and is widely regarded as the father of the field of biogeography. Dubbed by historian, Michael Flannery a "Victorian Indiana Jones," Wallace explored the Amazon, lived with headhunters, and even survived a shipwreck.
Yet one facet of Wallace's remarkable career has been ignored: His embrace of intelligent design.
The story of Wallace's doubts about the creative power of natural selection and his growing belief that nature requires an intelligent cause is one of the forgotten chapters in the history of science. But it's an intriguing story that sheds new light on current debates. In this DVD, take a fascinating journey with Wallace as he breaks with Darwin and comes to believe that nature is permeated with evidence of purposeful design.
(Full Runtime, including extra features, 54 minutes.)
The primary portion of this video, the 21-minute documentary section, streams live on YouTube, at https://www.youtube.com/watch?v=hxvAVln6HLI.
The DVD is available from Amazon for $6.61
This meeting WAS NOT be recorded or broadcast to a ZOOM audience.
February 17, 2024

MOVIE NIGHT We watched the 2018 movie by Creation Ministries International, "ALIEN INTRUSION -Unmasking a deception."
From the DVD cover: Million of people have seen UFO's and many even recall personal encounters with strange entities. The popular view is that these are advanced aliens visiting us from far, far away. Based upon the Amazon top-50, best-selling book, this documentary takes a deeper look at the events, the beliefs and the experts who have shaped our views in all things other worldly - including UFO sightings, Roswell, alien abductions and even alleged government coverups. And when one takes a deeper look at this phenomenon, it reveals one of the most disturbing, yet powerful affirmations of the truth of the Bible and Christianity. The truth is more surprising than most people realize.
(Runtime 109 minutes.)
The DVD is available from CMI for $12 plus shipping.
This meeting WILL NOT be recorded or broadcast to a ZOOM audience. To watch the video, you must attend in person.
From the DVD cover: Million of people have seen UFO's and many even recall personal encounters with strange entities. The popular view is that these are advanced aliens visiting us from far, far away. Based upon the Amazon top-50, best-selling book, this documentary takes a deeper look at the events, the beliefs and the experts who have shaped our views in all things other worldly - including UFO sightings, Roswell, alien abductions and even alleged government coverups. And when one takes a deeper look at this phenomenon, it reveals one of the most disturbing, yet powerful affirmations of the truth of the Bible and Christianity. The truth is more surprising than most people realize.
(Runtime 109 minutes.)
The DVD is available from CMI for $12 plus shipping.
This meeting WILL NOT be recorded or broadcast to a ZOOM audience. To watch the video, you must attend in person.
February 3, 2024

Russ Humphreys, Ph.D (Nuclear Physics) will gave a ZOOM presentation titled "A MORE BIBLICAL COSMOLOGY" from his home in Tennessee. This program was recorded. To watch, CLICK HERE: video4387396160.mp4.
About our speaker: (The following text was taken from the Creation Ministries International website)
Dr Humphreys was born on 2 February 1942 in Wyandotte, Michigan, U.S.A., and was raised in a scientifically aware but non–Christian household. Not surprisingly, Russell himself always had a love for science, and in 1959, he was one of the 40 winners of the Westinghouse National Science Talent Search.
He received a B.S. degree in physics at Duke University, 1959–1963. After this, he moved to Louisiana State University (LSU) to study postgraduate physics. In 1969, while doing his dissertation research for LSU in the mountains of Colorado, he committed his life to Christ. In 1972, he was awarded a Ph.D. in physics, on cosmic rays and ultrahigh energy nucleon–nucleon interactions, by which time he was a fully convinced creationist due to both the biblical and scientific evidence. For the next 6 years he worked in the High Voltage Laboratory of General Electric Company, designing and inventing equipment and researching high–voltage phenomena. While there, he received a U.S. patent and one of Industrial Research Magazine’s IR–100 awards.
Dr Humphreys has been married since 1963, and they have three children.
Research
Beginning in 1979 he worked for Sandia National Laboratories (New Mexico) in nuclear physics, geophysics, pulsed-power research, and theoretical atomic and nuclear physics. In 1985, he began working with Sandia’s ‘Particle Beam Fusion Project’, and was co-inventor of special laser-triggered ‘Rimfire’ high-voltage switches, now coming into wider use.
The last decade at Sandia saw greater emphasis on theoretical nuclear physics and radiation hydrodynamics in an effort to help produce the world’s first lab–scale thermonuclear fusion. Besides gaining two other U.S. patents, Dr Humphreys has been given two awards from Sandia, including an Award for Excellence for contributions to light ion–fusion target theory.
Overall, Dr Humphreys’ research has been very wide-ranging:
Creationist work full-time
Dr Humphreys retired from Sandia in 2001 to work full-time for the Institute for Creation Research (ICR) where he was appointed an Associate Professor of Physics. In this time, he operated mainly from his home office in Albuquerque, NM, USA, while still continuing to write for Journal of Creation (formerly TJ) and assisting several other creationist organizations with questions and information concerning physics, astronomy and cosmology.
In August 2008 he resigned from ICR to work for Creation Ministries International (USA) while continuing to be mostly located in New Mexico, continuing his research, plus writing and speaking ministry.
Dr Humphreys has been a longtime member of the Creation Research Society and is on its Board of Directors.
Education
B.S., Duke University, Durham, NC, 1963
Ph.D., Louisiana State University, Baton Rouge, LA, 1972
Honors/Awards/Associations
Creation Science Fellowship of New Mexico, board member and past President
Two of Industrial Research Magazine’s IR-100 awards
Award for Excellence for contributions to light ion-fusion target theory
Adjunct professor of the Institute for Creation Research in San Diego
Board member of the Creation Research Society
Member, American Geophysical Union
Dr. Humphreys is also a Logos Research Associate.
About our speaker: (The following text was taken from the Creation Ministries International website)
Dr Humphreys was born on 2 February 1942 in Wyandotte, Michigan, U.S.A., and was raised in a scientifically aware but non–Christian household. Not surprisingly, Russell himself always had a love for science, and in 1959, he was one of the 40 winners of the Westinghouse National Science Talent Search.
He received a B.S. degree in physics at Duke University, 1959–1963. After this, he moved to Louisiana State University (LSU) to study postgraduate physics. In 1969, while doing his dissertation research for LSU in the mountains of Colorado, he committed his life to Christ. In 1972, he was awarded a Ph.D. in physics, on cosmic rays and ultrahigh energy nucleon–nucleon interactions, by which time he was a fully convinced creationist due to both the biblical and scientific evidence. For the next 6 years he worked in the High Voltage Laboratory of General Electric Company, designing and inventing equipment and researching high–voltage phenomena. While there, he received a U.S. patent and one of Industrial Research Magazine’s IR–100 awards.
Dr Humphreys has been married since 1963, and they have three children.
Research
Beginning in 1979 he worked for Sandia National Laboratories (New Mexico) in nuclear physics, geophysics, pulsed-power research, and theoretical atomic and nuclear physics. In 1985, he began working with Sandia’s ‘Particle Beam Fusion Project’, and was co-inventor of special laser-triggered ‘Rimfire’ high-voltage switches, now coming into wider use.
The last decade at Sandia saw greater emphasis on theoretical nuclear physics and radiation hydrodynamics in an effort to help produce the world’s first lab–scale thermonuclear fusion. Besides gaining two other U.S. patents, Dr Humphreys has been given two awards from Sandia, including an Award for Excellence for contributions to light ion–fusion target theory.
Overall, Dr Humphreys’ research has been very wide-ranging:
- Designed and theoretically analyzed thermonuclear fusion targets using radiation hydrodynamic codes.
- Designed key high-voltage parts of Sandia’s 100-Terawatt Particle Beam Fusion Accelerator II and conducted fusion power experiments on it. Same designs are in use today on Sandia’s Z machine.
- Research on low-temperature solids and studies on superconductors.
- Nuclear weapons projects, including stockpile engineering for W87 firing set.
- Helped design new inkjet printer component and shared patent on it.
- Developed high repetition-rate neutron tube driver and gamma-ray spectrometer for borehole logging applications. Patent on high-voltage power supply for it.
- Patents on wide-bandwidth electric field sensor and high-voltage neutron tube supply.
- Designed lightning current waveform recorder which won IR-100 Award.
- Studied electric fields and ion currents under ultrahigh voltage DC transmission lines.
- Theoretical studies of relativistic velocity dependence of nuclear forces.
Creationist work full-time
Dr Humphreys retired from Sandia in 2001 to work full-time for the Institute for Creation Research (ICR) where he was appointed an Associate Professor of Physics. In this time, he operated mainly from his home office in Albuquerque, NM, USA, while still continuing to write for Journal of Creation (formerly TJ) and assisting several other creationist organizations with questions and information concerning physics, astronomy and cosmology.
In August 2008 he resigned from ICR to work for Creation Ministries International (USA) while continuing to be mostly located in New Mexico, continuing his research, plus writing and speaking ministry.
Dr Humphreys has been a longtime member of the Creation Research Society and is on its Board of Directors.
Education
B.S., Duke University, Durham, NC, 1963
Ph.D., Louisiana State University, Baton Rouge, LA, 1972
Honors/Awards/Associations
Creation Science Fellowship of New Mexico, board member and past President
Two of Industrial Research Magazine’s IR-100 awards
Award for Excellence for contributions to light ion-fusion target theory
Adjunct professor of the Institute for Creation Research in San Diego
Board member of the Creation Research Society
Member, American Geophysical Union
Dr. Humphreys is also a Logos Research Associate.
January 20, 2024

Jim Solliday, President of the Microscopy Society of Southern California, gave a presentation titled, FISH OF THE PALEOZOIC: TEETH & SCALES - Histology and morphology of Paleozoic fossils as revealed by high resolution light microscopy.
Surviving soft-tissues have been recovered and described from dinosaurs of the Cretaceous and now are widely accepted as endogenous. The question now is how far back in the geological record can preserved soft-tissues be found. Beyond the time of the dinosaurs lies the Paleozoic and supposedly a great stretch of time that includes the Permian to the Devonian. Evolution claims that during this time the first Tetrapods emerged from the water and walked on land. The Carboniferous and Devonian takes us back over 355 to 419 million years (by conventional wisdom), making it inconceivable that any remnant of original-tissue could persist and be revealed even under the best research microscopes. Are soft-tissue structures still present in Paleozoic fossils?
This study attempts to answers that question. We suspect that if most of the creatures died during the great Flood the answer may be, yes. The overall scope is to investigate the potential confirmation of vertebrate Soft-Tissues all the way back to the beginning. To provide a more complete description of vertebrate fossils from Paleozoic deposits one needs to extend the investigation to the microscopic level. The primary goal is to evaluate the preservation and information potential of this rare and unique biota. This investigation has been restricted to mainly a histological and morphological study. Imaging with high-resolution microscopy has been directed to the cell level. More advanced elemental analysis is beyond the scope of this study and reserved for future consideration. The source material is primarily from, but not limited to the lower main coal shale of Northumberland and Durham coal-fields. This is geographically located in the north-east regains of England. From these lower coal measures a valuable collection of fish and reptiles has been established (Lower Carboniferous). Histological sections are employed for characterizing the micro-morphology. Some of the thin-sections were originally from the collection of the British 1980’s TV botanist David Bellamy. This study provides a tour of the rare micro vertebrate fossils found in Carboniferous deposits (Paleozoic) from the North of England.
Research by. James Solliday, Pres. Microscopical Society of Southern California, EPL
About our speaker:
Jim Solliday was the 2019 recipient of the State Microscopical Society of Illinois (SMSI) 's August Köhler Award, the most prestigious honor given in the field of microscopy. SMSI published the following recognition of Jim's many accomplishments:
James D. Solliday is recognized for his numerous contributions to light microscopy and active participation in microscopical societies. He was a principal founder of the Microscopical Society of Southern California (MSSC) in 1996 and is currently the MSSC president, a chair he has held every year since 2002. He is also a long-time contributor to the Los Angeles Microscopical Society, and a member of the American Circuit of the Postal Microscopical Society, Quekett Microscopical Club, and the International Society for Diatom Research. Mr. Solliday is well known for his extensive knowledge of microscope history. He assists microscope collectors worldwide and was the curator of the Frank Lundy Antique Microscope Collection, which was publicly exhibited at the San Francisco international airport.
Mr. Solliday has taught workshops on diatoms, illumination methods, staining, photomicrography, microscope maintenance, and the history and identification of antique microscopes. He has published dozens of microscopy articles, including contributions to the MSSC, San Francisco Microscopical Society, Los Angeles Microscopical Society, and The Quekett Journal of Microscopy. He was instrumental in obtaining and donating surplus microscopes to area schools, and has been a regular visiting lecturer to local classrooms.
For 35 years, Mr. Solliday has operated his own business called Educational Photo Lab, which specializes in photomicrography. He has produced photomicrographs for Hollywood movies, documentaries, litigation exhibits, contamination analysis, authors, Ph.D. students, and publishers, with more than 450 microscope images published in textbooks. Beyond the micro world, he served for 31 years as a professional firefighter and medical technician for the city of Costa Mesa, CA.
Surviving soft-tissues have been recovered and described from dinosaurs of the Cretaceous and now are widely accepted as endogenous. The question now is how far back in the geological record can preserved soft-tissues be found. Beyond the time of the dinosaurs lies the Paleozoic and supposedly a great stretch of time that includes the Permian to the Devonian. Evolution claims that during this time the first Tetrapods emerged from the water and walked on land. The Carboniferous and Devonian takes us back over 355 to 419 million years (by conventional wisdom), making it inconceivable that any remnant of original-tissue could persist and be revealed even under the best research microscopes. Are soft-tissue structures still present in Paleozoic fossils?
This study attempts to answers that question. We suspect that if most of the creatures died during the great Flood the answer may be, yes. The overall scope is to investigate the potential confirmation of vertebrate Soft-Tissues all the way back to the beginning. To provide a more complete description of vertebrate fossils from Paleozoic deposits one needs to extend the investigation to the microscopic level. The primary goal is to evaluate the preservation and information potential of this rare and unique biota. This investigation has been restricted to mainly a histological and morphological study. Imaging with high-resolution microscopy has been directed to the cell level. More advanced elemental analysis is beyond the scope of this study and reserved for future consideration. The source material is primarily from, but not limited to the lower main coal shale of Northumberland and Durham coal-fields. This is geographically located in the north-east regains of England. From these lower coal measures a valuable collection of fish and reptiles has been established (Lower Carboniferous). Histological sections are employed for characterizing the micro-morphology. Some of the thin-sections were originally from the collection of the British 1980’s TV botanist David Bellamy. This study provides a tour of the rare micro vertebrate fossils found in Carboniferous deposits (Paleozoic) from the North of England.
Research by. James Solliday, Pres. Microscopical Society of Southern California, EPL
About our speaker:
Jim Solliday was the 2019 recipient of the State Microscopical Society of Illinois (SMSI) 's August Köhler Award, the most prestigious honor given in the field of microscopy. SMSI published the following recognition of Jim's many accomplishments:
James D. Solliday is recognized for his numerous contributions to light microscopy and active participation in microscopical societies. He was a principal founder of the Microscopical Society of Southern California (MSSC) in 1996 and is currently the MSSC president, a chair he has held every year since 2002. He is also a long-time contributor to the Los Angeles Microscopical Society, and a member of the American Circuit of the Postal Microscopical Society, Quekett Microscopical Club, and the International Society for Diatom Research. Mr. Solliday is well known for his extensive knowledge of microscope history. He assists microscope collectors worldwide and was the curator of the Frank Lundy Antique Microscope Collection, which was publicly exhibited at the San Francisco international airport.
Mr. Solliday has taught workshops on diatoms, illumination methods, staining, photomicrography, microscope maintenance, and the history and identification of antique microscopes. He has published dozens of microscopy articles, including contributions to the MSSC, San Francisco Microscopical Society, Los Angeles Microscopical Society, and The Quekett Journal of Microscopy. He was instrumental in obtaining and donating surplus microscopes to area schools, and has been a regular visiting lecturer to local classrooms.
For 35 years, Mr. Solliday has operated his own business called Educational Photo Lab, which specializes in photomicrography. He has produced photomicrographs for Hollywood movies, documentaries, litigation exhibits, contamination analysis, authors, Ph.D. students, and publishers, with more than 450 microscope images published in textbooks. Beyond the micro world, he served for 31 years as a professional firefighter and medical technician for the city of Costa Mesa, CA.
January 6, 2024

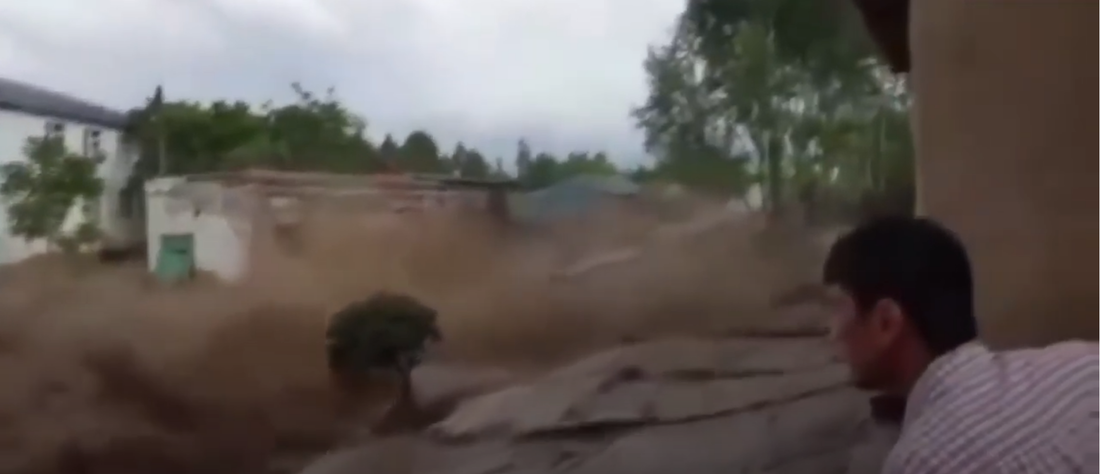
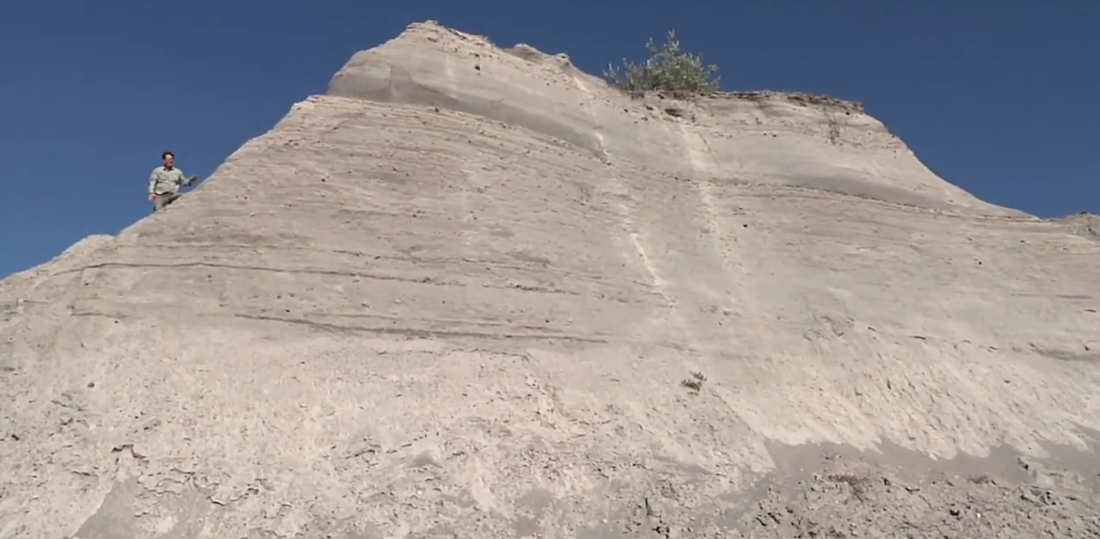
MOVIE NIGHT: We watched the newly released Is Genesis History? sequel, MOUNTAINS AFTER THE FLOOD.
IN THIS SEQUEL to Is Genesis History? Dr. Del Tacket follows a team of creation scientists as they uncover amazing new evidence for the global Flood. You'll see enormous folded rock layers, study geological maps, watch experiments in a laboratory, and fly over the Grand Canyon. By the time the journey is over, you'll have experienced a completely new perspective on what happened at the end of the Flood. From microscopes to mountaintops, from ancient lakes to modern laboratories, watch creation scientists do their incredible work of discovery.
IN THIS SEQUEL to Is Genesis History? Dr. Del Tacket follows a team of creation scientists as they uncover amazing new evidence for the global Flood. You'll see enormous folded rock layers, study geological maps, watch experiments in a laboratory, and fly over the Grand Canyon. By the time the journey is over, you'll have experienced a completely new perspective on what happened at the end of the Flood. From microscopes to mountaintops, from ancient lakes to modern laboratories, watch creation scientists do their incredible work of discovery.
December 2 & 16, 2023

Calvary Chapel Central Orange County honors the Lord's birth with a flurry of special events throughout the month of December. To accommodate the church's extra busy schedule during Christmas Month, the Creation Science Fellowship did not meet in December.
November 18, 2023

Joseph Kezele, MD, will gave a LIVE, IN-PERSON presentation titled, GOD'S AMAZING CREATURES #3.
(This presentation was recorded, and will be viewable at this location sin the near future. Please check back.)
About our speaker: Joseph Kezele studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomat of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians.
He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, and is Associate Professor of Biology at Arizona Christian University in Glendale, teaching Nutrition and Wellness, Medical terminology, Biochemistry, Pathophysiology, Organic Chemistry and Creation Science Apologetics.
Dr. Kezele is also a Logos Research Associate, and had the privilege of participating in a dinosaur dig in Montana where he excavated a triceratops horn and the triceratops cervical condyle from which intact osteocytes in soft tissue were isolated.
He has participated in 24 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, Armenia and Moldova. Dr. Kezele is one of the principal presenters in the Institute for Creation Research’s 4 DVD series Made in His Image. Dr. Kezele teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona, where he and his wife Patty attend. Their three adult children live and work in the Phoenix area.
(This presentation was recorded, and will be viewable at this location sin the near future. Please check back.)
About our speaker: Joseph Kezele studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomat of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians.
He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, and is Associate Professor of Biology at Arizona Christian University in Glendale, teaching Nutrition and Wellness, Medical terminology, Biochemistry, Pathophysiology, Organic Chemistry and Creation Science Apologetics.
Dr. Kezele is also a Logos Research Associate, and had the privilege of participating in a dinosaur dig in Montana where he excavated a triceratops horn and the triceratops cervical condyle from which intact osteocytes in soft tissue were isolated.
He has participated in 24 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, Armenia and Moldova. Dr. Kezele is one of the principal presenters in the Institute for Creation Research’s 4 DVD series Made in His Image. Dr. Kezele teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona, where he and his wife Patty attend. Their three adult children live and work in the Phoenix area.
November 4, 2023
|
DVD available from Answers in Genesis Bookstore.
|
We watched the AIG dvd, "REPLACING DARWIN," featuring Nathaniel Jeanson, Ph.D. (Cell and developmental biology) From the DVD jacket: "Science has come a long way since Darwin made his claims in the 1800s. Several remarkable discoveries shed new light on the earth's origins. In addition, much of the data Darwin presented for his ideas no longer seem like clear evidence for evolution. Dr. Nathaniel Jeanson effectively and authoritatively documents the newest scientific discoveries, refutes Darwin's claims, and presents evidences that support a compelling alternative explanation for the origin of species." .............................................................. Dr. Nathaniel T. Jeanson earned his BS in molecular biology and bioinformatics from the University of Wisconsin-Parkside and his Ph.D. from Harvard University. His research findings have been presented at national and regional conferences and have been published in peer-reviewed journals, such as Blood, Nature, and Cell. Since 2009, he has been actively researching the origin of species, both at the Institute for Creation Research and at Answers in Genesis. ......................................................... This link will take you to a YouTube video of Dr. Jeanson giving a presentation of the same title at a different venue. |
October 21, 2023

Ross Anderson, Ph.D. (Molecular Biology) will give* an in-house presentation titled, THE FORMATION OF ANTIBODIES.
With all the Covid stuff behind us more people have become familiar with antibodies, but how many really know what an antibody molecule is, how they are formed and what triggers their release? A brief look at some of the intricacies of the immune system reveal design at the highest most complex levels. We will briefly take a closer look into the extreme complexity and design of just one small but very important aspect of the immune system.
Also, we have invited Christine Anderson, Dr. Ross's lovely wife, to perform a handbell-ringing concert for our edification.
About our speaker:
Dr. Anderson is newly retired from The Masters University. Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society. Dr. Anderson is also a Logos Research Associate.
With all the Covid stuff behind us more people have become familiar with antibodies, but how many really know what an antibody molecule is, how they are formed and what triggers their release? A brief look at some of the intricacies of the immune system reveal design at the highest most complex levels. We will briefly take a closer look into the extreme complexity and design of just one small but very important aspect of the immune system.
Also, we have invited Christine Anderson, Dr. Ross's lovely wife, to perform a handbell-ringing concert for our edification.
About our speaker:
Dr. Anderson is newly retired from The Masters University. Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society. Dr. Anderson is also a Logos Research Associate.

About our concert bell-ringer (from the website, Voicesinbronze.com):
The solo handbell artistry of Christine D. Anderson is world-renowned. With finesse, grace and dexterity, Christine's solo handbell artistry has thrilled audiences in 25 countries and 50 states, and her fans around the world will tell you that you'll never be the same after hearing and seeing Christine play.
Christine began her handbell ringing career at Coral Ridge Presbyterian Church, Ft. Lauderdale, Florida, in late 1972. After completing a B.A. in music at Florida Atlantic University, she directed 6 handbell choirs at Coral Ridge and began conducting workshops and festivals around the country.
Christine rang her first handbell solo, "The Lord’s Prayer," in 1980, discovering the gift that would define her life’s work and produced a video called "Voices in Bronze®." She writes for several music journals, is on the editorial board for “Creator Magazine”, and serves the Southern Baptist Convention as Editor for "Handbells" magazine. Christine has published over 120 solo handbell arrangements, all reflecting her commitment to excellence in handbell ringing. She has been an annual recipient of the ASCAP Standards Award for compositions and performances for many years.
The solo handbell artistry of Christine D. Anderson is world-renowned. With finesse, grace and dexterity, Christine's solo handbell artistry has thrilled audiences in 25 countries and 50 states, and her fans around the world will tell you that you'll never be the same after hearing and seeing Christine play.
Christine began her handbell ringing career at Coral Ridge Presbyterian Church, Ft. Lauderdale, Florida, in late 1972. After completing a B.A. in music at Florida Atlantic University, she directed 6 handbell choirs at Coral Ridge and began conducting workshops and festivals around the country.
Christine rang her first handbell solo, "The Lord’s Prayer," in 1980, discovering the gift that would define her life’s work and produced a video called "Voices in Bronze®." She writes for several music journals, is on the editorial board for “Creator Magazine”, and serves the Southern Baptist Convention as Editor for "Handbells" magazine. Christine has published over 120 solo handbell arrangements, all reflecting her commitment to excellence in handbell ringing. She has been an annual recipient of the ASCAP Standards Award for compositions and performances for many years.
october 7, 2023

Greg Morgan, retired Nuclear Safety Engineer with the U.S. Department of Energy (DOE), gave an in-house presentation titled, FLOOD CURRENTS FROZEN IN STONE.
Greg Morgan writes: Does it matter whether the Earth is young? It mattered to me. I could not accept salvation until I had a reason to believe that I needed a savior.
Greg Morgan writes: Does it matter whether the Earth is young? It mattered to me. I could not accept salvation until I had a reason to believe that I needed a savior.
September 16, 2023

Tim Standish, Ph.D. (Environmental Biology and Public Policy) gave a presentation titled "SIGNS OF DESIGN"
Abstract:
Philosophers struggle with what evidence might be a sign of design, but normal people don’t have that problem. It would take a lot of “education” to become blinded to the fact that a Boeing 787 aircraft, or even the worst book ever written, was designed. The same is true of the signs of design we see in life. We will look at some evidence from recent papers about living thing, you can decide whether or not there are signs of design, but be sure to ask yourself if your evaluation is a result of the evidence and logic, or something else.
About our speaker:
Dr. Standish is Adjunct Associate Professor of Earth and Biological Sciences in the Loma Linda University's School of Medicine. He holds a B.S. in zoology from Andrews University, an M.S. in biology from Andrews University, and a Ph.D. in biology and public policy from George Mason University (University of Virginia), Charlottesville, Virginia.
SIGNS OF DESIGN PowerPoint presentation
Recommendation: CLICK to open the first slide below, then progress by CLICKING arrows at far right.
Abstract:
Philosophers struggle with what evidence might be a sign of design, but normal people don’t have that problem. It would take a lot of “education” to become blinded to the fact that a Boeing 787 aircraft, or even the worst book ever written, was designed. The same is true of the signs of design we see in life. We will look at some evidence from recent papers about living thing, you can decide whether or not there are signs of design, but be sure to ask yourself if your evaluation is a result of the evidence and logic, or something else.
About our speaker:
Dr. Standish is Adjunct Associate Professor of Earth and Biological Sciences in the Loma Linda University's School of Medicine. He holds a B.S. in zoology from Andrews University, an M.S. in biology from Andrews University, and a Ph.D. in biology and public policy from George Mason University (University of Virginia), Charlottesville, Virginia.
SIGNS OF DESIGN PowerPoint presentation
Recommendation: CLICK to open the first slide below, then progress by CLICKING arrows at far right.
September 2, 2023

Heinz Lycklama, Ph.D. (Nuclear Physics), gave a ZOOM presentation titled, "A RETROSPECTIVE ON THE COVID PHENOMENON."
https://heinzlycklama.com/videos/Presentations/CovidPhenomenonCSF-090223.mp4
A Retrospective On The COVID Phenomenon: The Covid-19 phenomenon has been with us since March 2020 and has had a very disruptive impact on the world. We review Covid tests, cases, and deaths, and the impact of the use of masks, vaccines, mandates and lockdowns. We have learned much about the use of therapeutics and vaccines in treating Covid-19 over the last two years. In particular, we examine what the V-Safe system, the VAERS data, and VAERS safety signals are are telling us. The Sudden Adult Death Syndrome, excess deaths, and economic impacts are raising many concerns. What should we as believers do with what we have learned from our experience with Covid-19? Acts 19:32, Acts 17:11, 1 Th. 5:21, 2 Tim. 1:7.
The PowerPoint slides Dr. Lycklama used for this presentation may be found here: www.heinzlycklama.com/messages/csf-23/COVID-Retrospective.pdf
About our ZOOM speaker:
Summary of Professional Career in High Tech Industry (borrowed from CreationWiki's biography for Dr. Lycklama):
https://heinzlycklama.com/videos/Presentations/CovidPhenomenonCSF-090223.mp4
A Retrospective On The COVID Phenomenon: The Covid-19 phenomenon has been with us since March 2020 and has had a very disruptive impact on the world. We review Covid tests, cases, and deaths, and the impact of the use of masks, vaccines, mandates and lockdowns. We have learned much about the use of therapeutics and vaccines in treating Covid-19 over the last two years. In particular, we examine what the V-Safe system, the VAERS data, and VAERS safety signals are are telling us. The Sudden Adult Death Syndrome, excess deaths, and economic impacts are raising many concerns. What should we as believers do with what we have learned from our experience with Covid-19? Acts 19:32, Acts 17:11, 1 Th. 5:21, 2 Tim. 1:7.
The PowerPoint slides Dr. Lycklama used for this presentation may be found here: www.heinzlycklama.com/messages/csf-23/COVID-Retrospective.pdf
About our ZOOM speaker:
Summary of Professional Career in High Tech Industry (borrowed from CreationWiki's biography for Dr. Lycklama):
- 1969-1977, Member of Technical Staff, Bell Telephone Laboratories in Murray Hill, NJ
- 1977-78, Supervisor, Bell Telephone Laboratories in Holmdel, NJ
- 1978-1986, Vice President Development, INTERACTIVE Systems Corp. in Santa Monica, CA
- 1986-1991, Senior Vice President of Technology, INTERACTIVE Systems Corp. in Santa Monica, CA
- 1992- 2013, President of Open Systems Technology Associates (OSTA) in CA and WA
- 2013 – present, Apologetics Forum Founder, Author, Speaker, Consultant.
August 19, 2023

Thomas Kindell, Ph.D. gave a presentation* called BIOMEMETICS - Part 6.
https://drive.google.com/file/d/1jahD7MiyMOxa4ACW1HdtdWgX2QdyOx4l/view
About our speaker (borrowed from Creation Wiki):
Dr. Thomas Kindell was once an ardent believer in the “fact” of evolution. However, through his exposure to the scientific case for creation, he became a zealous creationist. He has received advanced training in scientific creationism through the Graduate School of the Institute for Creation Research in Santee, California. He has been privileged to study under several of the world's most prominent creation scientists. He also has studied Christian Apologetics and Biblical-Scientific Creationism at California Graduate School of Theology where he received his M.A. in Biblical Studies. He holds a Doctorate in Philosophy of Theology (major in philosophy of Biblical apologetics.)
Dr. Kindell is the founder and president of Reasons for Faith Ministries, Inc., a non-profit ministry dedicated to equipping Christian believers to “give every man an answer” for their Biblical faith. He is a licensed Assemblies of God evangelist and is the only credentialed Assemblies minister devoted to full-time teaching of Creation-Science apologetics. He is a member of the Oregon Design Science Association and is a lifetime sustaining member of the Creation Research Society.
For over twenty-eight years he has engaged in a traveling teaching ministry lecturing on Biblical apologetics and the scientific case for creation. Dr. Kindell is available for speaking engagements full-time on a free will offering basis throughout the United States and abroad. His seminars have been enthusiastically endorsed by Dr. Duane T. Gish, Senior Vice‑President Emeritus of the Institute for Creation Research. He has also successfully debated the scientific case for creation against notable evolutionists on high school and college campuses, radio, and television. Dr. Kindell is a frequent guest on the radio series “Science, Scripture & Salvation” which is produced by the Institute for Creation Research. He is the author of the acclaimed book, Evolution on Trial.
https://drive.google.com/file/d/1jahD7MiyMOxa4ACW1HdtdWgX2QdyOx4l/view
About our speaker (borrowed from Creation Wiki):
Dr. Thomas Kindell was once an ardent believer in the “fact” of evolution. However, through his exposure to the scientific case for creation, he became a zealous creationist. He has received advanced training in scientific creationism through the Graduate School of the Institute for Creation Research in Santee, California. He has been privileged to study under several of the world's most prominent creation scientists. He also has studied Christian Apologetics and Biblical-Scientific Creationism at California Graduate School of Theology where he received his M.A. in Biblical Studies. He holds a Doctorate in Philosophy of Theology (major in philosophy of Biblical apologetics.)
Dr. Kindell is the founder and president of Reasons for Faith Ministries, Inc., a non-profit ministry dedicated to equipping Christian believers to “give every man an answer” for their Biblical faith. He is a licensed Assemblies of God evangelist and is the only credentialed Assemblies minister devoted to full-time teaching of Creation-Science apologetics. He is a member of the Oregon Design Science Association and is a lifetime sustaining member of the Creation Research Society.
For over twenty-eight years he has engaged in a traveling teaching ministry lecturing on Biblical apologetics and the scientific case for creation. Dr. Kindell is available for speaking engagements full-time on a free will offering basis throughout the United States and abroad. His seminars have been enthusiastically endorsed by Dr. Duane T. Gish, Senior Vice‑President Emeritus of the Institute for Creation Research. He has also successfully debated the scientific case for creation against notable evolutionists on high school and college campuses, radio, and television. Dr. Kindell is a frequent guest on the radio series “Science, Scripture & Salvation” which is produced by the Institute for Creation Research. He is the author of the acclaimed book, Evolution on Trial.
August 5, 2023

We watched a Logos Research Associates video, titled, MONKEY BUSINESS IN THE CHIMP GENOME by Jeffrey Tomkins, Ph.D. ( Genetics )
Evolutionists continue to contend that genetic studies have proved humans and chimpanzees share a common ancestor and have a 98-99% genome similarity. But many people don’t realize the current chimpanzee genome has not been constructed on its own merits. When genomes (a complete set of chromosomes for a given creature) are sequenced, the initial DNA is obtained in very small pieces and then assembled—essentially “stitched” together—with the aid of a computer. Since the chimpanzee genome hadn’t been previously sequenced, it lacked a good genetic framework to help guide the process. Given a strong evolutionary bias that humans evolved from a chimp-like ancestor, how do you suppose scientists assembled the chimpanzee DNA sequences? You guessed it! They used the human genome as a guide.
But there’s even more monkey business involved in producing the chimp genome. Human DNA contamination is also present and may have produced a flawed chimp genome that would appear to be far more human-like than it is.
Hear Dr Tomkins discuss his research project investigating these issues and the results of his comparison of over 2.5 million raw chimpanzee DNA sequences to both the human genome and the current version of the chimp genome. These results clearly show that many regions of the chimp genome are misassembled, the genetic similarity of the chimp genome to the human genome is much less than 98-99% and therefore can’t be used to support human evolution.
This video is available to watch at your convenience at https://www.youtube.com/watch?v=zPUe3sm3PvI
1:23 Running time..
Evolutionists continue to contend that genetic studies have proved humans and chimpanzees share a common ancestor and have a 98-99% genome similarity. But many people don’t realize the current chimpanzee genome has not been constructed on its own merits. When genomes (a complete set of chromosomes for a given creature) are sequenced, the initial DNA is obtained in very small pieces and then assembled—essentially “stitched” together—with the aid of a computer. Since the chimpanzee genome hadn’t been previously sequenced, it lacked a good genetic framework to help guide the process. Given a strong evolutionary bias that humans evolved from a chimp-like ancestor, how do you suppose scientists assembled the chimpanzee DNA sequences? You guessed it! They used the human genome as a guide.
But there’s even more monkey business involved in producing the chimp genome. Human DNA contamination is also present and may have produced a flawed chimp genome that would appear to be far more human-like than it is.
Hear Dr Tomkins discuss his research project investigating these issues and the results of his comparison of over 2.5 million raw chimpanzee DNA sequences to both the human genome and the current version of the chimp genome. These results clearly show that many regions of the chimp genome are misassembled, the genetic similarity of the chimp genome to the human genome is much less than 98-99% and therefore can’t be used to support human evolution.
This video is available to watch at your convenience at https://www.youtube.com/watch?v=zPUe3sm3PvI
1:23 Running time..

About our video speaker:
Dr. Tomkins earned a master’s degree in plant science in 1990 from the University of Idaho, where he performed research in plant hormones. He received his Ph.D. in genetics from Clemson University in 1996. While at Clemson, he worked as a research technician in a plant breeding/genetics program, with a research focus in the area of quantitative and physiological genetics in soybean. After receiving his Ph.D., he worked at a genomics institute and became a faculty member in the Department of Genetics and Biochemistry at Clemson. He had become a Christian as an undergraduate at Washington State University in 1982, with a goal to eventually work as a scientist and author in the creation science field. In 2009, Dr. Tomkins joined the Institute for Creation Research. He is the primary author of The Design and Complexity of the Cell and Chimps and Humans: A Geneticist Discovers DNA Evidence That Challenges Evolution, a co-author of Fascinating Creatures: Evidence of Christ's Handiwork and Human Origins, and a contributor to Guide to Creation Basics and Creation Basics & Beyond.
Dr. Tomkins is a Research Scientist at the Institute for Creation Research (ICR), and a Logos Research Associate with Logos Research Associates, (LogosRA).
Dr. Tomkins earned a master’s degree in plant science in 1990 from the University of Idaho, where he performed research in plant hormones. He received his Ph.D. in genetics from Clemson University in 1996. While at Clemson, he worked as a research technician in a plant breeding/genetics program, with a research focus in the area of quantitative and physiological genetics in soybean. After receiving his Ph.D., he worked at a genomics institute and became a faculty member in the Department of Genetics and Biochemistry at Clemson. He had become a Christian as an undergraduate at Washington State University in 1982, with a goal to eventually work as a scientist and author in the creation science field. In 2009, Dr. Tomkins joined the Institute for Creation Research. He is the primary author of The Design and Complexity of the Cell and Chimps and Humans: A Geneticist Discovers DNA Evidence That Challenges Evolution, a co-author of Fascinating Creatures: Evidence of Christ's Handiwork and Human Origins, and a contributor to Guide to Creation Basics and Creation Basics & Beyond.
Dr. Tomkins is a Research Scientist at the Institute for Creation Research (ICR), and a Logos Research Associate with Logos Research Associates, (LogosRA).
July 15, 2023

We watched a Logos Research Associates video titled, FOSSIL PROTEINS FROM FOUR CONTINENTS with Brian Thomas, Ph.D. (Paleobiochemistry).
Were red blood cells really discovered inside dinosaur bones? Are fossilized structures that look just like blood vessels, cells, and skin actually from the original creatures? YES! Original organic materials show why rocks and fossils look thousands not millions of years old. Dr. Thomas will show that original biochemistry is geologically extensive, geographically global, and taxonomically wide-ranging!
You can watch this video at your convenience at https://www.youtube.com/watch?v=mBPAlR41nTE
1:19 Running time
About our speaker:
Dr. Brian Thomas, Research Scientist specializing in Dinosaurs, Problems with Evolution, Human OriginsBrian Thomas received a master’s in biotechnology in 1999 from Stephen F. Austin State University, Nacogdoches, Texas, and a Ph.D. in paleobiochemistry in 2019 from the University of Liverpool. He taught junior high and high school at Christian schools in Texas, as well as biology, chemistry, and anatomy as an adjunct and assistant professor at Dallas-area universities. In 2008 Dr. Thomas joined the Institute for Creation Research as a science writer and editor, contributing news and magazine articles, speaking on creation issues, and researching original tissue fossils. He was appointed as Research Associate in 2019 and Research Scientist in 2021. He is the author of Dinosaurs and the Bible; co-author of Parks Across America: Viewing God's Wonders Through a Creationist Lens, Fascinating Creatures: Evidence of Christ's Handiwork, and Human Origins; and a contributor to Guide to Creation Basics, Creation Basics & Beyond, Guide to Dinosaurs, Guide to the Human Body, Guide to the Universe, and Dinosaurs: God’s Mysterious Creatures. His dissertation, Ancient and Fossil Bone Collagen Remnants, is available in book form.
Were red blood cells really discovered inside dinosaur bones? Are fossilized structures that look just like blood vessels, cells, and skin actually from the original creatures? YES! Original organic materials show why rocks and fossils look thousands not millions of years old. Dr. Thomas will show that original biochemistry is geologically extensive, geographically global, and taxonomically wide-ranging!
You can watch this video at your convenience at https://www.youtube.com/watch?v=mBPAlR41nTE
1:19 Running time
About our speaker:
Dr. Brian Thomas, Research Scientist specializing in Dinosaurs, Problems with Evolution, Human OriginsBrian Thomas received a master’s in biotechnology in 1999 from Stephen F. Austin State University, Nacogdoches, Texas, and a Ph.D. in paleobiochemistry in 2019 from the University of Liverpool. He taught junior high and high school at Christian schools in Texas, as well as biology, chemistry, and anatomy as an adjunct and assistant professor at Dallas-area universities. In 2008 Dr. Thomas joined the Institute for Creation Research as a science writer and editor, contributing news and magazine articles, speaking on creation issues, and researching original tissue fossils. He was appointed as Research Associate in 2019 and Research Scientist in 2021. He is the author of Dinosaurs and the Bible; co-author of Parks Across America: Viewing God's Wonders Through a Creationist Lens, Fascinating Creatures: Evidence of Christ's Handiwork, and Human Origins; and a contributor to Guide to Creation Basics, Creation Basics & Beyond, Guide to Dinosaurs, Guide to the Human Body, Guide to the Universe, and Dinosaurs: God’s Mysterious Creatures. His dissertation, Ancient and Fossil Bone Collagen Remnants, is available in book form.
july 1, 2023

We watched the videos, PANDEMIC COUP D' ETAT', PARTS 1&2
Why? The COVID-19 Pandemic is a creation science issue - a Frankenstein concoction served up by human creators who played god at the the world's peril.
Why? The COVID-19 Pandemic is a creation science issue - a Frankenstein concoction served up by human creators who played god at the the world's peril.
June 17, 2023

Joseph Kezele, MD, will give a LIVE, IN-PERSON presentation titled, Does Sickle Cell Anemia Provide Evidence for Evolution? (50 minutes) .Begins with looking at the production and structure of hemoglobin, the material that carries oxygen in the blood. Then the role of the mutation responsible for sickle cell anemia is examined, followed by the question of sickle cell anemia providing protection against malaria. The toll of misery and suffering of sickle cell anemia is presented.
About our speaker: Joseph Kezele studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomat of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians.
He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, and is Associate Professor of Biology at Arizona Christian University in Glendale, teaching Nutrition and Wellness, Medical terminology, Biochemistry, Pathophysiology, Organic Chemistry and Creation Science Apologetics.
Dr. Kezele is also a Logos Research Associate, and had the privilege of participating in a dinosaur dig in Montana and excavated a triceratops horn and the triceratops cervical condyle from which intact osteocytes in soft tissue were isolated.
He has participated in 24 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, Armenia and Moldova. Dr. Kezele is one of the principal presenters in the Institute for Creation Research’s 4 DVD series Made in His Image. Dr. Kezele teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona, where he and his wife Patty attend. Their three adult children live and work in the Phoenix area.
About our speaker: Joseph Kezele studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomat of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians.
He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, and is Associate Professor of Biology at Arizona Christian University in Glendale, teaching Nutrition and Wellness, Medical terminology, Biochemistry, Pathophysiology, Organic Chemistry and Creation Science Apologetics.
Dr. Kezele is also a Logos Research Associate, and had the privilege of participating in a dinosaur dig in Montana and excavated a triceratops horn and the triceratops cervical condyle from which intact osteocytes in soft tissue were isolated.
He has participated in 24 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, Armenia and Moldova. Dr. Kezele is one of the principal presenters in the Institute for Creation Research’s 4 DVD series Made in His Image. Dr. Kezele teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona, where he and his wife Patty attend. Their three adult children live and work in the Phoenix area.
JUNE 3, 2023

SUSPENDED ANIMATION - is it in the Bible?
In the sense that death prevents animation until the resurrection, yes.
In the sense of suspended animation being a state of existence somewhere between life and death, probably not - though that condition possibly could be lurking in the background behind a some passages.
We discussed those passages, and watched a fascinating video about advancements in the technology of suspended animation for humans.
In the sense that death prevents animation until the resurrection, yes.
In the sense of suspended animation being a state of existence somewhere between life and death, probably not - though that condition possibly could be lurking in the background behind a some passages.
We discussed those passages, and watched a fascinating video about advancements in the technology of suspended animation for humans.
may 20, 2023

ARMING Students against Evolutionary Indoctrination
We took a quick survey of FREE online videos and other resources for enabling students to refute evolutionary propaganda.
This included 30 one-minute (or less) vignettes of informed students politely challenging evolutionary claims made in the classroom, from Genesis Apologetics at
https://genesisapologetics.com/classroom/ . It also included video links at Answers in Genesis (AIG), https://answersingenesis.org/media/ ; The Institute for Creation Research (ICR), https://www.icr.org/thatsafact/ ; Creation Ministries International (CMI), https://www.youtube.com/playlist?list=PLtDG1Hzii2vtPqO-wMV7hETspC5rZDISn ;Logos Research Associates (LogosRA), https://www.youtube.com/@logosresearchassociates; and
Beyond Is Genesis History, https://isgenesishistory.com/tag/video/
We took a quick survey of FREE online videos and other resources for enabling students to refute evolutionary propaganda.
This included 30 one-minute (or less) vignettes of informed students politely challenging evolutionary claims made in the classroom, from Genesis Apologetics at
https://genesisapologetics.com/classroom/ . It also included video links at Answers in Genesis (AIG), https://answersingenesis.org/media/ ; The Institute for Creation Research (ICR), https://www.icr.org/thatsafact/ ; Creation Ministries International (CMI), https://www.youtube.com/playlist?list=PLtDG1Hzii2vtPqO-wMV7hETspC5rZDISn ;Logos Research Associates (LogosRA), https://www.youtube.com/@logosresearchassociates; and
Beyond Is Genesis History, https://isgenesishistory.com/tag/video/
May 6, 2023

We watched video recordings of Chemist, James Tour, Ph,D., - a latter day Bezaleel (see Exodus 31:1-5 below), Raised as a secular Jew, James encountered Messiah while in college, and has sought daily to serve and honor Him ever since. It has pleased the LORD to pour wisdom, understanding, knowledge, and skill into Dr. Tour across multiple disciplines, appointing him head of three different departments at Rice University, and blessing him and his students with many groundbreaking discoveries and inventions. These promise to change technology across the world on multiple fronts, including medicine, energy, nanotechnology, materials engineering, construction, etc!
About our featured video speaker (biography borrowed from the Discovery Institute website, https://www.discovery.org/p/tour/ )
JAMES TOUR PROFESSOR OF CHEMISTRY, OF COMPUTER SCIENCE, AND OF MATERIALS SCIENCE AND NANO-ENGINEERING
James Tour is the T. T. and W. F. Chao Professor of Chemistry, Professor of Computer Science, and Professor of Materials Science and Nano-Engineering at Rice University. A synthetic organic chemist, he received his BS in Chemistry from Syracuse University, his PhD in synthetic organic and organometallic chemistry from Purdue University, and postdoctoral training in synthetic organic chemistry at the University of Wisconsin and Stanford University. He has served on the faculty of the University of South Carolina and as a visiting scholar at Harvard University.
Tour has over 700 research publications and over 130 patent families.
He was inducted into the National Academy of Inventors in 2015 and was listed in “The World’s Most Influential Scientific Minds” by Thomson Reuters in 2014. He has been named “Scientist of the Year” by R&D Magazine and was ranked one of the Top 10 chemists in the world over the past decade by a Thomson Reuters citations per publication index survey in 2009. The same year he was elected a Fellow of the American Association for the Advancement of Science (AAAS). He won the Feynman Prize in Experimental Nanotechnology in 2008, the NASA Space Act Award in 2008 for his development of carbon nanotube reinforced elastomers, and the Arthur C. Cope Scholar Award from the American Chemical Society for his achievements in organic chemistry in 2007.
# # #
Exodus 31:1-5
And the LORD spake unto Moses, saying, See, I have called by name Bezaleel the son of Uri, the son of Hur, of the tribe of Judah:
And I have filled him with the spirit of God, in wisdom, and in understanding, and in knowledge, and in all manner of workmanship, To devise cunning works, to work in gold, and in silver, and in brass, And in cutting of stones, to set them, and in carving of timber, to work in all manner of workmanship.
For those who cannot join us in person or online for this program, the videos we plan to watch are as follow:
https://www.youtube.com/watch?v=kqz-dCIUa8Y
Jim Garlow and James Tour discuss new breakthroughs in chemistry (36min)
* * *
https://www.youtube.com/watch?v=OKPGIdpPeS4
Saving the Planet with the Nanotech Revolution! Turning Trash into Cash (27min)
* * *
https://www.youtube.com/watch?v=gm4DWnD72TE
Resurrection: Can the Bible be Trusted? | Dr. James Tour (59min)
About our featured video speaker (biography borrowed from the Discovery Institute website, https://www.discovery.org/p/tour/ )
JAMES TOUR PROFESSOR OF CHEMISTRY, OF COMPUTER SCIENCE, AND OF MATERIALS SCIENCE AND NANO-ENGINEERING
James Tour is the T. T. and W. F. Chao Professor of Chemistry, Professor of Computer Science, and Professor of Materials Science and Nano-Engineering at Rice University. A synthetic organic chemist, he received his BS in Chemistry from Syracuse University, his PhD in synthetic organic and organometallic chemistry from Purdue University, and postdoctoral training in synthetic organic chemistry at the University of Wisconsin and Stanford University. He has served on the faculty of the University of South Carolina and as a visiting scholar at Harvard University.
Tour has over 700 research publications and over 130 patent families.
He was inducted into the National Academy of Inventors in 2015 and was listed in “The World’s Most Influential Scientific Minds” by Thomson Reuters in 2014. He has been named “Scientist of the Year” by R&D Magazine and was ranked one of the Top 10 chemists in the world over the past decade by a Thomson Reuters citations per publication index survey in 2009. The same year he was elected a Fellow of the American Association for the Advancement of Science (AAAS). He won the Feynman Prize in Experimental Nanotechnology in 2008, the NASA Space Act Award in 2008 for his development of carbon nanotube reinforced elastomers, and the Arthur C. Cope Scholar Award from the American Chemical Society for his achievements in organic chemistry in 2007.
# # #
Exodus 31:1-5
And the LORD spake unto Moses, saying, See, I have called by name Bezaleel the son of Uri, the son of Hur, of the tribe of Judah:
And I have filled him with the spirit of God, in wisdom, and in understanding, and in knowledge, and in all manner of workmanship, To devise cunning works, to work in gold, and in silver, and in brass, And in cutting of stones, to set them, and in carving of timber, to work in all manner of workmanship.
For those who cannot join us in person or online for this program, the videos we plan to watch are as follow:
https://www.youtube.com/watch?v=kqz-dCIUa8Y
Jim Garlow and James Tour discuss new breakthroughs in chemistry (36min)
* * *
https://www.youtube.com/watch?v=OKPGIdpPeS4
Saving the Planet with the Nanotech Revolution! Turning Trash into Cash (27min)
* * *
https://www.youtube.com/watch?v=gm4DWnD72TE
Resurrection: Can the Bible be Trusted? | Dr. James Tour (59min)
april 15, 2023

Heinz Lycklama, Ph.D. (Nuclear Physics), will gave a presentation titled, "Answering The New Atheists."
Some “New Atheists” have written hard-hitting books over the last decade claiming that God does not exist and that religion poisons everything. They (Dawkins, Dennett, Harris, Hitchens) are often referred to as the “Four Horsemen.” Dr. Lycklama looked at the principles of the new atheism and considered eight of their common claims. A number of Christian Apologists (Wilson, Fernandes, McDurmon, Lennox) have written books refuting the claims of these atheists. Dr. Lycklama showed that the atheist’s claims are nothing new and are indeed false. In doing so he looked at a wide range of evidence and asked the question – does the evidence best support Theism or Atheism?
Summary of Professional Career in High Tech Industry (borrowed from CreationWiki's biography for Dr. Lycklama):
Some “New Atheists” have written hard-hitting books over the last decade claiming that God does not exist and that religion poisons everything. They (Dawkins, Dennett, Harris, Hitchens) are often referred to as the “Four Horsemen.” Dr. Lycklama looked at the principles of the new atheism and considered eight of their common claims. A number of Christian Apologists (Wilson, Fernandes, McDurmon, Lennox) have written books refuting the claims of these atheists. Dr. Lycklama showed that the atheist’s claims are nothing new and are indeed false. In doing so he looked at a wide range of evidence and asked the question – does the evidence best support Theism or Atheism?
Summary of Professional Career in High Tech Industry (borrowed from CreationWiki's biography for Dr. Lycklama):
- 1969-1977, Member of Technical Staff, Bell Telephone Laboratories in Murray Hill, NJ
- 1977-78, Supervisor, Bell Telephone Laboratories in Holmdel, NJ
- 1978-1986, Vice President Development, INTERACTIVE Systems Corp. in Santa Monica, CA
- 1986-1991, Senior Vice President of Technology, INTERACTIVE Systems Corp. in Santa Monica, CA
- 1992- 2013, President of Open Systems Technology Associates (OSTA) in CA and WA
- 2013 – present, Apologetics Forum Founder, Author, Speaker, Consultant.
april 1, 2023
We watched Part 2 of the GENESIS CONFERENCE that was held at Jessup University, March 4, 2023

LINEUP:
1) Dr. Randy Guliuzza
2) David Rives
3) Dr. Dan Biddle
4) Panel Discussion of Speakers
You may watch the conference by CLICKING here:
(https://genesisapologetics.com/g1-conference/)
1) Dr. Randy Guliuzza
2) David Rives
3) Dr. Dan Biddle
4) Panel Discussion of Speakers
You may watch the conference by CLICKING here:
(https://genesisapologetics.com/g1-conference/)
march 18, 2023

LINEUP:
1) Dr. John Jackson
2) Dr. Dan Biddle
3) Pastor Joe Walters
4) Dr. Randy Guliuzza
5) David Rives
You may watch the conference by CLICKING here:
(https://genesisapologetics.com/g1-conference/)
1) Dr. John Jackson
2) Dr. Dan Biddle
3) Pastor Joe Walters
4) Dr. Randy Guliuzza
5) David Rives
You may watch the conference by CLICKING here:
(https://genesisapologetics.com/g1-conference/)
march 4, 2023

We watched the video, DIED SUDDENLY 2022 (FULL DOCUMENTARY)
Broadly speaking, Creation Science encompasses more than the genius and splendor of God's Creation, but also such wretchedly dangerous human creations as the COVID-19 "VACCINES".
You can access this video at https://rumble.com/v1wcs7o-died-suddenly-2022-full-documentary.html
Broadly speaking, Creation Science encompasses more than the genius and splendor of God's Creation, but also such wretchedly dangerous human creations as the COVID-19 "VACCINES".
You can access this video at https://rumble.com/v1wcs7o-died-suddenly-2022-full-documentary.html
Warning: The contents of this presentation are not suitable for children.
Warning: The contents of this presentation are not suitable for children.
February 18, 2023
We watched (1) the 45 minute Koinonia House video, ISSACHAR INSIGHT, followed by two 30 minute videos from the Genesis Science Research ANOMALIES series titled, (2) SPEED OF LIGHT and (3) ZERO POINT ENERGY.
(1) ISSACHAR INSIGHT - Chuck Missler and Barry Setterfield. The late Chuck Missler interviews Barry Setterfield about Zero Point Energy, the speed of light, and plasma physics at large in the Universe. [Backup link: ISSACHAR INSIGHT - Chuck Missler and Barry Setterfield]
Genesis Science Research (2) LIGHTSPEED A Journey of Discovery
Genesis Science Research (2) LIGHTSPEED A Journey of Discovery
Is the Velocity of Light Constant in Time? by Alan Montgomery, mathematician, and Lambert Dolphin, physicistA Determination and Analysis of Appropriate Values of the Speed of Light to Test the Setterfield Hypothesis by Alan Montgomery, mathematician
FEBRUARY 4, 2023

We watched the video, PLASMA ASTRONOMY AND THE BIBLE, followed by a live ZOOM Q&A with the author, Barry Setterfield of Speed of Light Decay fame.
For more than a hundred years, the studies of astronomy and cosmology have been dominated by gravitational physics, with little thought given to the influences of electric and magnetic fields on a cosmic scale. This is surprising, because:
For more than a hundred years, the studies of astronomy and cosmology have been dominated by gravitational physics, with little thought given to the influences of electric and magnetic fields on a cosmic scale. This is surprising, because:
- 99.9% of the universe is electrified gas, either partially or fully ionized PLASMA - the fourth state of matter.
- Gravitational force is far, far weaker than electric and magnetic forces.
- Observations of stars and galaxies on an astronomic scale mirror observations of plasma behavior on a laboratory scale.
- It stands to reason that electric and magnetic fields and currents should play a major role in the shaping of stars and galaxies.
About our speaker: Barry Setterfield was born in Australia and educated at the University of Adelaide. Mr. Setterfield is an active researcher who has been investigating the speed of light since 1979. His first major paper was published by Flinders University in 1987, co-authored by Trevor Norman, and written at the request of a senior physicist at Stanford Research Institute International.
Research has continued in the fields of physics, geology, and astronomy, with papers published in the Journal of Theoretics. He and his wife are currently living in southern Oregon, where he recently retired from the position of Director of the New Hope Observatory. (For a more complete biography, click here: https://barrysetterfield.org/GSRbiography.html)
Research has continued in the fields of physics, geology, and astronomy, with papers published in the Journal of Theoretics. He and his wife are currently living in southern Oregon, where he recently retired from the position of Director of the New Hope Observatory. (For a more complete biography, click here: https://barrysetterfield.org/GSRbiography.html)
january 21, 2023

John R. Baumgardner, Ph.D. Geophysics and Space Physics, gave a presentation, titled
RATE REVISITED: RADIOISOTOPE EVIDENCE THAT THE EARTH IS YOUNG
Nuclear decay of uranium generates stable daughter elements lead and helium. The Radioisotopes and the Age of the Earth (RATE) project (1997-2005) carefully measured the rate at which helium migrates through the mineral zircon. These measurements showed that at temperatures typical of the Earth’s surface, even tiny fractions of the helium generated by uranium decay cannot be retained for millions or billions of years. Yet zircons from a 4 km deep drill core into the granite crust in New Mexico were found to contain up to 80% of the helium produced from 1.5 billion years’ worth of uranium to lead decay—at presently observed decay rates—still present inside them. This was one of three prominent lines of radioisotope evidence studied by the RATE team that nuclear decay rates have not been constant during earth history but instead have been orders of magnitude higher during brief episodes in the past.
A second powerful line of evidence that pointed to this same conclusion involved microscopic spherical patterns of radiation damage in rocks known as a radio halos. While radio halos produced by U-238 decay are well understood, radio halos produced by the element polonium are not. Since their discovery early in the 20 th century, polonium radio halos have been viewed as an unexplained enigma within the scientific world. Two major reasons are (1) the half-lives of polonium isotopes are so short—3 minutes,64 micro-seconds, and 138 days for Po-218, Po-214, and Po-210 and (2) the halos contain no tiny crystal at their center able to host the original polonium. The RATE team isolated and photographed a total of31,000 radiohalos from 50 granite formations around the world. Of this total, 19,000 were formed exclusively by polonium decay. The geological context in which these Po halos occur, mostly in the cores of cooling granite magma bodies, imply that they were formed in a short span of time. This research also requires a brief episode of exceedingly rapid nuclear decay. Episodes of extremely high rates of nuclear decay documented by the RATE team imply that (1) dates provided by standard radioisotope methods are in error by large factors and (2) the true age of the Earth is only a few thousands of years instead of billions.
RATE REVISITED: RADIOISOTOPE EVIDENCE THAT THE EARTH IS YOUNG
Nuclear decay of uranium generates stable daughter elements lead and helium. The Radioisotopes and the Age of the Earth (RATE) project (1997-2005) carefully measured the rate at which helium migrates through the mineral zircon. These measurements showed that at temperatures typical of the Earth’s surface, even tiny fractions of the helium generated by uranium decay cannot be retained for millions or billions of years. Yet zircons from a 4 km deep drill core into the granite crust in New Mexico were found to contain up to 80% of the helium produced from 1.5 billion years’ worth of uranium to lead decay—at presently observed decay rates—still present inside them. This was one of three prominent lines of radioisotope evidence studied by the RATE team that nuclear decay rates have not been constant during earth history but instead have been orders of magnitude higher during brief episodes in the past.
A second powerful line of evidence that pointed to this same conclusion involved microscopic spherical patterns of radiation damage in rocks known as a radio halos. While radio halos produced by U-238 decay are well understood, radio halos produced by the element polonium are not. Since their discovery early in the 20 th century, polonium radio halos have been viewed as an unexplained enigma within the scientific world. Two major reasons are (1) the half-lives of polonium isotopes are so short—3 minutes,64 micro-seconds, and 138 days for Po-218, Po-214, and Po-210 and (2) the halos contain no tiny crystal at their center able to host the original polonium. The RATE team isolated and photographed a total of31,000 radiohalos from 50 granite formations around the world. Of this total, 19,000 were formed exclusively by polonium decay. The geological context in which these Po halos occur, mostly in the cores of cooling granite magma bodies, imply that they were formed in a short span of time. This research also requires a brief episode of exceedingly rapid nuclear decay. Episodes of extremely high rates of nuclear decay documented by the RATE team imply that (1) dates provided by standard radioisotope methods are in error by large factors and (2) the true age of the Earth is only a few thousands of years instead of billions.
january 7, 2023
movie night

Critics challenge the Bible's teaching that God created the heavens and the earth only six thousand years ago because it conflicts with conventional wisdom that (a) the speed of light is universal, and (b) the radius of the visible Universe is nearly 14 billion lightyears. How could the Universe be only 6,000 years old if light from distant galaxies has been in route to us at constant velocity for many billions of years?
(By the way, secular cosmologists have their own light-time problem, one for which they invented an "inflationary period," in which the universe underwent an extremely rapid exponential expansion, MANY times faster than the speed of light).
One proposed solution to the "Distant Starlight Problem" is the idea that the speed of light may vary with direction. Tonight, we watched excerpts from Dr. Jason Lisle's "The Distant Starlight Solution," Dr. Derek Muller's Veritasium video, "Why No One Has Measured the Speed of Light," the Slow Mo Guys, Gaven Free's and Daniel Gruchy's video, "Filming the Speed of Light at 10 Trillion FPS," and "Determining the speed of light at home using chocolate and a microwave and how Louis Essen did it," by Paul Looyen with High School Physics Explained.
A "convention" that the speed of light is infinite in one direction, but only c/2 upon reflection appears to be ruled out on two counts: (1) Pulses of light recored at 10 trillion frames per second. bouncing around inside multi-faceted mirrored chamber, give no indication of changing their speed with change in direction. (2) Nodes of wave interference in microwave ovens (resonant chambers) should not exist if light velocity, (and hence wavelength), were infinite in one direction.
(By the way, secular cosmologists have their own light-time problem, one for which they invented an "inflationary period," in which the universe underwent an extremely rapid exponential expansion, MANY times faster than the speed of light).
One proposed solution to the "Distant Starlight Problem" is the idea that the speed of light may vary with direction. Tonight, we watched excerpts from Dr. Jason Lisle's "The Distant Starlight Solution," Dr. Derek Muller's Veritasium video, "Why No One Has Measured the Speed of Light," the Slow Mo Guys, Gaven Free's and Daniel Gruchy's video, "Filming the Speed of Light at 10 Trillion FPS," and "Determining the speed of light at home using chocolate and a microwave and how Louis Essen did it," by Paul Looyen with High School Physics Explained.
A "convention" that the speed of light is infinite in one direction, but only c/2 upon reflection appears to be ruled out on two counts: (1) Pulses of light recored at 10 trillion frames per second. bouncing around inside multi-faceted mirrored chamber, give no indication of changing their speed with change in direction. (2) Nodes of wave interference in microwave ovens (resonant chambers) should not exist if light velocity, (and hence wavelength), were infinite in one direction.
December 17, 2022

Calvary Chapel Tustin honors the Lord's birth with a flurry of special events and activities throughout the month of December. In order to accommodate these extra activities, the Creation Science Fellowship has agreed to skip what would normally be its second meeting of the month meeting on December 17, and resume regular programming on the first Saturday of the new year, January 7, 2023.
december 3, 2022

Jerry Bergman, Ph.D. (Human Biology); Ph.D. (Measurement and Evaluation); M.S.O.H. (Occupational Health); M.P.H. M.P.H. (Public Health); M.S.B.S. (Biomedical Science); M.A. (Social Psychology); M.Ed. (Counseling and psychology); B.S., (Sociology, Biology, Psychology); A.A. (Biology, Behavioral Science) gave a presentation, titled,
"Darwin Demolished: Why evolution is unequivocally Impossible.". Settled science documents the main pillars of evolution have been demolished. The question why evolution is widely believed when it goes against settled science is answered. The presentation is documented by 50 stunning illustrations.
About our speaker:
(The following excerpt from another website -creation.com/dr-jerry-bergmanwebsite - is antiquated and in need of substantial updating. Dr. Bergman now has well over 100 publications to his name, including numerous books he has authored since the time this excerpt was posted.):
Jerry Bergman has taught biology, genetics, chemistry, biochemistry, anthropology, geology, and microbiology at Northwest State College in Archbold OH for over 25 years. He has 9 degrees, including 7 graduate (= ‘post-graduate’ in some non-US systems) degrees. Dr Bergman is a graduate of Medical College of Ohio, Wayne State University in Detroit, The University of Toledo, and Bowling Green State University. He has over 800 publications in 12 languages and 20 books and monographs. He has also taught at the Medical College of Ohio where he was a research associate in the department of experimental pathology, and he also taught 6 years at the University of Toledo, and 7 years at Bowling Green State University.
Among his books is a monograph on peer evaluation published by the College Student Journal Press, a Fastback on the creation-evolution controversy published by Phi Delta Kappa, a book on vestigial organs with Dr George Howe (‘Vestigial Organs’ are Fully Functional), a book on psychology and religious cults, a book on religious discrimination published by Onesimus Press, and a book on mental health published by Claudius Verlag in München. He has also published a college textbook on evaluation (Boston, Houghton Mifflin Co.), and has contributed to dozens of other textbooks. He was also a consultant for over 20 science text books, mostly biology and biochemistry.
Dr Bergman has presented over one hundred scientific papers at professional and community meetings in the United States, Canada, and Europe. To discuss his research, he has been a featured speaker on many college campuses throughout the United States and Europe, and is a frequent guest on radio and television programs. His research has made the front page in newspapers throughout the country, has been featured by the Paul Harvey Show several times, and has been discussed by David Brinkley, Chuck Colson, and other nationally known commentators on national television.
His other work experience includes over ten years experience at various Mental Health/Psychology clinics as a licensed professional clinical counselor and three years full time corrections research for a large county circuit court in Michigan and inside the walls of Jackson Prison (SPSM), the largest walled prison in the world. He has also served as a consultant for CBS News, ABC News, Reader’s Digest, Amnesty International, several government agencies and for two Nobel Prize winners, including the inventor of the transistor. In the past decade he has consulted or has testified as an expert witness or consultant in almost one-hundred court cases. A Fellow of the American Scientific Association, member of The National Association for the Advancement of Science, and many other professional associations, he is listed in Who’s Who in America, Who’s Who in the Midwest and in Who’s Who in Science and Religion.
Education
ArticlesHis articles on creation include:
"Darwin Demolished: Why evolution is unequivocally Impossible.". Settled science documents the main pillars of evolution have been demolished. The question why evolution is widely believed when it goes against settled science is answered. The presentation is documented by 50 stunning illustrations.
About our speaker:
(The following excerpt from another website -creation.com/dr-jerry-bergmanwebsite - is antiquated and in need of substantial updating. Dr. Bergman now has well over 100 publications to his name, including numerous books he has authored since the time this excerpt was posted.):
Jerry Bergman has taught biology, genetics, chemistry, biochemistry, anthropology, geology, and microbiology at Northwest State College in Archbold OH for over 25 years. He has 9 degrees, including 7 graduate (= ‘post-graduate’ in some non-US systems) degrees. Dr Bergman is a graduate of Medical College of Ohio, Wayne State University in Detroit, The University of Toledo, and Bowling Green State University. He has over 800 publications in 12 languages and 20 books and monographs. He has also taught at the Medical College of Ohio where he was a research associate in the department of experimental pathology, and he also taught 6 years at the University of Toledo, and 7 years at Bowling Green State University.
Among his books is a monograph on peer evaluation published by the College Student Journal Press, a Fastback on the creation-evolution controversy published by Phi Delta Kappa, a book on vestigial organs with Dr George Howe (‘Vestigial Organs’ are Fully Functional), a book on psychology and religious cults, a book on religious discrimination published by Onesimus Press, and a book on mental health published by Claudius Verlag in München. He has also published a college textbook on evaluation (Boston, Houghton Mifflin Co.), and has contributed to dozens of other textbooks. He was also a consultant for over 20 science text books, mostly biology and biochemistry.
Dr Bergman has presented over one hundred scientific papers at professional and community meetings in the United States, Canada, and Europe. To discuss his research, he has been a featured speaker on many college campuses throughout the United States and Europe, and is a frequent guest on radio and television programs. His research has made the front page in newspapers throughout the country, has been featured by the Paul Harvey Show several times, and has been discussed by David Brinkley, Chuck Colson, and other nationally known commentators on national television.
His other work experience includes over ten years experience at various Mental Health/Psychology clinics as a licensed professional clinical counselor and three years full time corrections research for a large county circuit court in Michigan and inside the walls of Jackson Prison (SPSM), the largest walled prison in the world. He has also served as a consultant for CBS News, ABC News, Reader’s Digest, Amnesty International, several government agencies and for two Nobel Prize winners, including the inventor of the transistor. In the past decade he has consulted or has testified as an expert witness or consultant in almost one-hundred court cases. A Fellow of the American Scientific Association, member of The National Association for the Advancement of Science, and many other professional associations, he is listed in Who’s Who in America, Who’s Who in the Midwest and in Who’s Who in Science and Religion.
Education
- M.S.O.H. (Master of Science in Occupational Health), Medical College of Ohio, Toledo, Ohio, 2004.
- M.P.H. M.P.H. (Master of Science in Public Health), Northwest Ohio Consortium for Public Health (Medical College of Ohio, Toledo, Ohio; University of Toledo, Toledo, Ohio; Bowling Green State University, Bowling Green, Ohio), 2001.
- M.S.B.S. (Master of Science in Biomedical Science), Medical College of Ohio, Toledo, Ohio, 1999.
- Ph.D. in human biology, Columbia Pacific University, San Rafael, California, 1992.
- M.A. in social psychology, Bowling Green State University, Bowling Green, Ohio, 1986.
- Ph.D. in measurement and evaluation, minor in psychology, Wayne State University, Detroit, Michigan, 1976.
- M.Ed. in counseling and psychology, Wayne State University, Detroit, Michigan, 1971.
- B.S., Wayne State University, Detroit, Michigan, 1970. Major area of study was sociology, biology, and psychology.
- A.A. in Biology and Behavioral Science, Oakland Community College, Bloomfield Hills, Michigan, 1967.
- Fellow of the American Scientific Affiliation, 1983
- Who’s Who in America
- MENSA
- Ohio certification to teach both elementary and high school levels
- National Association for Gifted Children
- American Educational Research Association
- National Council on Measurement in Education
- American Sociological Association
- American Psychological Association
- Ohio Psychological Association
- Association for the Scientific Study of Religion
- American Association of Suicidology
- Institute of Religion and Health
- American Society of Corrections
- Ohio Science Teachers Association.
- American Biology Teachers Association.
- The American Scientific Affiliation.
- The American Association for the Advancement of Science.
- The American Association for the History of Science
- American Chemical Society
- American Institute of Biological Sciences.
- Ohio Academy of Science
- American Institute of Chemists
- New York Academy of Sciences
- The New York Museum of Natural History
- Society for the Scientific Study of Male Psychology and Physiology, President and Founder
ArticlesHis articles on creation include:
- Research has overturned endosymbiosis: the unbridgeable gap between prokaryotes and eukaryotes remains
- The central role of Darwinism in the Holocaust
- New fossil proves turtle evolution … or does it?
- Academia and the press as the bad guys—review of Spectacle: The astonishing life of Ota Benga by Pamela Newkirk
- Darwinists still trying to refute Behe and still failing—review of Darwin Devolves: The new science about DNA that challenges evolution by Michael Behe
- Common examples of ‘one gene, one trait’ exposed
- Fingernails and toenails—useless evolutionary relics or an important part of design?
- A book about human errors that don’t exist—review of Human Errors: A panorama of our glitches, from pointless bones to broken genes by Nathan H. Lents
- A history of the United Methodist Church’s opposition to creationism and intelligent design
- Darwin’s Point
- Developmental gene regulatory networks—an insurmountable impediment to evolution
- Neutral Model, genetic drift and the Third Way—a synopsis of the self-inflicted demise of the evolutionary paradigm
- Is the male reproductive system poorly designed?
- The ‘common sense’ argument for evolution
- Arthur I. Brown: An early creation leader
- Dino-bird theory—a flight of fancy
- Darwin’s corrosive influence on literature and society as a whole. A review of Apostate—The Men Who Destroyed the Christian West by Kevin Swanson
- Marcel-Paul Schützenberger—French Darwin doubter
- Feathers—an evolutionary enigma
- C. Everett Koop—Christian and Darwin doubter
- James Tour—leading scientist and Darwin skeptic
- Incomplete lineage sorting and other ‘rogue’ data fell the tree of life
- The left recurrent laryngeal nerve design in mammals is not poor design
- The sea horse
- Did Darwin plagiarize his evolution theory?
- The tragic toll of toxic teaching: A review of Saving Darwin: How to be a Christian and Believe in Evolution by Karl W. Giberson
- Is the human genome nearly identical to chimpanzee?—a reassessment of the literature
- Genomic monkey business—estimates of nearly identical human–chimp DNA similarity re-evaluated using omitted data
- The chromosome 2 fusion model of human evolution—part 1: re-evaluating the evidence
- The chromosome 2 fusion model of human evolution—part 2: re-analysis of the genomic data
- Darwin is the universal acid that affects everything
- From skepticism to faith in Christ: a Nobel Laureate’s journey
- Anders Breivik—Social Darwinism leads to mass murder
- Using facial angle to prove evolution and the human race hierarchy
- John C. Eccles, Nobel laureate and Darwin doubter
- Robert A. Millikan, physics Noble laureate and Darwin doubter
- Chapter from In Six Days: Why 50 Scientists Choose to Believe in Creation
- Are wisdom teeth (third molars) vestiges of human evolution?
- The Dodo Bird … An example of survival of the fittest
- Who invented it first?
- Do any vestigial organs exist in humans?
- Ancon sheep: just another loss mutation
- The evolution of feathers: a major problem for Darwinism
- Understanding Poisons from a Creationist Perspective
- Darwinism and the Nazi race Holocaust
- H.G. Wells: Darwin’s disciple and eugenicist extraordinaire
- The church preaches eugenics: a history of church support for Darwinism and eugenics: review of Preaching Genetics: Religious Leaders and the American Eugenics Movement by Christine Rosen
- The Darwinian foundation of communism
- The miracle of tears
- Why the Miller–Urey research argues against abiogenesis
- Creationism and the problem of homosexual behavior
- Teaching Creation and Evolution in Public Schools Solid research reveals American beliefs
- Did God Make Pathogenic Viruses?
- The history of the teaching of human female inferiority in Darwinism
- Evolutionary naturalism: an ancient idea
- ATP: The Perfect Energy Currency for the Cell (CRSQ article posted on True Origins website)
- Why Abiogenesis is Impossible (CRSQ article posted on True Origins website)
- Ota Benga: the pygmy put on display in a zoo
- Is the human male nipple vestigial?
- The design of tears: an example of irreducible complexity
- Darwin’s critical influence on the ruthless extremes of capitalism
- Is the human pharynx poorly designed?
- The flat-earth myth and creationism
- Did immune system antibody diversity evolve?
- Birth control leader Margaret Sanger: Darwinist, racist and eugenicist
- Does homology provide evidence of evolutionary naturalism?
- Ludwig Wittgenstein: Darwin doubter
november 19, 2022

John R. Baumgardner, Ph.D. Geophysics and Space Physics, gave a presentation, titled
THE ROLE OF LARGE TSUNAMIS IN THE GENERATION OF THE FLOOD SEDIMENT RECORD.
This presentation explored the role of large tsunamis in the production of the fossil-bearing sediment record during the Genesis Flood. The study utilizes the Mabbul code that models the erosion, suspension, transport, and deposition of sediment by turbulent water on a rotating sphere. It assumes that repetitive, large-amplitude tsunamis occurred on segments of oceanic subduction zones, provided
the energy for driving the water motion and that cavitation was the primary erosion mechanism. This study includes a plate motion history for the various continental blocks spanning, in terms of geological nomenclature, the Paleozoic, Mesozoic, and Cenozoic eras, that is, the portion of the geological record formed during the Flood. That history begins with a supercontinent configuration known in the secular literature known as Pannotia which breaks apart and then reassembles as Pangea. That configuration in turn breaks apart and the resulting blocks migrate to the produce the continent configuration we have today. Most of the erosion takes place on the edges of the continent blocks. The resulting sediment is carried inland by the tsunamis as they sweep across the continental surfaces. The total sediment
produced approaches the amount on earth today. The spatial distribution of sediment matches that of the earth today to an astonishing degree. The large number of flat-topped seamounts, known as guyots, in the oceans today testify strongly to the reality of ferocious tsunamis that were taking place during the Flood cataclysm. This work seems to go a long way toward accounting for today’s sediment record in terms of the Genesis Flood only a few thousand years ago.
Due to technical difficulties, we were unable to record this lecture on November 19. However, Dr. Baumgardner spoke on the same topic at an earlier date to the Midwest Creation Fellowship. Here is that presentation. https://www.youtube.com/watch?v=AasMA5A21Tk
THE ROLE OF LARGE TSUNAMIS IN THE GENERATION OF THE FLOOD SEDIMENT RECORD.
This presentation explored the role of large tsunamis in the production of the fossil-bearing sediment record during the Genesis Flood. The study utilizes the Mabbul code that models the erosion, suspension, transport, and deposition of sediment by turbulent water on a rotating sphere. It assumes that repetitive, large-amplitude tsunamis occurred on segments of oceanic subduction zones, provided
the energy for driving the water motion and that cavitation was the primary erosion mechanism. This study includes a plate motion history for the various continental blocks spanning, in terms of geological nomenclature, the Paleozoic, Mesozoic, and Cenozoic eras, that is, the portion of the geological record formed during the Flood. That history begins with a supercontinent configuration known in the secular literature known as Pannotia which breaks apart and then reassembles as Pangea. That configuration in turn breaks apart and the resulting blocks migrate to the produce the continent configuration we have today. Most of the erosion takes place on the edges of the continent blocks. The resulting sediment is carried inland by the tsunamis as they sweep across the continental surfaces. The total sediment
produced approaches the amount on earth today. The spatial distribution of sediment matches that of the earth today to an astonishing degree. The large number of flat-topped seamounts, known as guyots, in the oceans today testify strongly to the reality of ferocious tsunamis that were taking place during the Flood cataclysm. This work seems to go a long way toward accounting for today’s sediment record in terms of the Genesis Flood only a few thousand years ago.
Due to technical difficulties, we were unable to record this lecture on November 19. However, Dr. Baumgardner spoke on the same topic at an earlier date to the Midwest Creation Fellowship. Here is that presentation. https://www.youtube.com/watch?v=AasMA5A21Tk
november 16, 2022
A SPECIALLY SCHEDULED 7:00 p.m. Wednesday night meeting of the Creation Science Fellowship -

Mark Armitage, Ed.D., with the Dinosaur Soft Tissue Research Institute (dstri.org) gave a LIVE report on his ongoing research in Dinosaur Soft Tissue!
NOTE: This report included the little known fact that many dinosaur fossils include un-mineralized remains containing identifiable cells and tissues!
This meeting was for serious students of all ages, grade school through senior citizens! Families were welcomed.
While supplies lasted, those in attendance will received a FREE 25-30 page book and an actual sample of un-mineralized dinosaur bone TO KEEP!
NOTE: This report included the little known fact that many dinosaur fossils include un-mineralized remains containing identifiable cells and tissues!
This meeting was for serious students of all ages, grade school through senior citizens! Families were welcomed.
While supplies lasted, those in attendance will received a FREE 25-30 page book and an actual sample of un-mineralized dinosaur bone TO KEEP!
november 5, 2022
The November 5 meeting was cancelled due to unforeseen circumstances.
October 1, 2022

Ross Anderson, Ph.D. Biology, gave a LIVE, IN-PERSON presentation, "EVIDENCE OF FORETHOUGHT AND DESIGN FROM BIOCHEMISTRY, MOLECULAR BIOLOGY, AND CELL BIOLOGY."
The living cell is replete with examples of forethought and design. Dr. Anderson gave examples from three different areas. We looked at the very complex molecular machine responsible for synthesizing ATP, the high-energy molecules required to power life. We will also examined protein sorting by the cell; How does the cell “know” which proteins go where? We will also learned one of the many ways in which a cell regulates which proteins to make and when to make them.
You may watch the recorded presentation here.
september 17, 2022

John Baumgardner, Ph.D., Geophysics and Space Physics, gave a remote presentation by ZOOM titled, GEOLOGICAL OBSERVATION POWERFULLY AFFIRMS THE REALITY OF THE GLOBAL FLOOD OF SCRIPTURE.
The vast lateral extent of sedimentary strata and a general lack of erosional channeling across layer boundaries point to global scale processes in the past radically different from the erosion, transport, and deposition processes occurring on Earth today. To deposit such uniform, laterally extensive layers with flat boundaries between them implies that vast portions of the continental surface were flooded by rapidly moving water and were nearly smooth. The water speed had to be high to keep sediment particles in suspension over such large distances. Further, fossilization generally requires that an organism be buried quickly and entirely, which is close to synonymous with catastrophic conditions. The ubiquity of fossils across the Phanerozoic portion of the geological record argues that this entire portion of the record is the product of catastrophe. These and other features of the record are consistent with the cataclysm described in Genesis chapters 7 and 8. This presentation will survey these observations and highlight the evidence that the cataclysm involved very rapid plate tectonics and rapid movement of
rock within the Earth’s mantle. It will also present the basics of the mechanism responsible for this cataclysm that involves a runaway instability in the mantle that reduced the shear strength of mantle minerals by a factor of a billion, led to a rapid mantle overturn, generated the entirety of today’s igneous ocean floor, and caused continental blocks to migrate thousands of miles across the face of the earth, all within a span of only a few months’ time. The still cool rock from the pre-Flood ocean floor now lies at the bottom of the lower mantle on its boundary with the outer core and is a prominent feature in all current seismic tomography images of the mantle. This mechanism is generally referred to as catastrophic plate tectonics.
About our speaker: (Please see the August 6 review below.)
The vast lateral extent of sedimentary strata and a general lack of erosional channeling across layer boundaries point to global scale processes in the past radically different from the erosion, transport, and deposition processes occurring on Earth today. To deposit such uniform, laterally extensive layers with flat boundaries between them implies that vast portions of the continental surface were flooded by rapidly moving water and were nearly smooth. The water speed had to be high to keep sediment particles in suspension over such large distances. Further, fossilization generally requires that an organism be buried quickly and entirely, which is close to synonymous with catastrophic conditions. The ubiquity of fossils across the Phanerozoic portion of the geological record argues that this entire portion of the record is the product of catastrophe. These and other features of the record are consistent with the cataclysm described in Genesis chapters 7 and 8. This presentation will survey these observations and highlight the evidence that the cataclysm involved very rapid plate tectonics and rapid movement of
rock within the Earth’s mantle. It will also present the basics of the mechanism responsible for this cataclysm that involves a runaway instability in the mantle that reduced the shear strength of mantle minerals by a factor of a billion, led to a rapid mantle overturn, generated the entirety of today’s igneous ocean floor, and caused continental blocks to migrate thousands of miles across the face of the earth, all within a span of only a few months’ time. The still cool rock from the pre-Flood ocean floor now lies at the bottom of the lower mantle on its boundary with the outer core and is a prominent feature in all current seismic tomography images of the mantle. This mechanism is generally referred to as catastrophic plate tectonics.
About our speaker: (Please see the August 6 review below.)
September 3, 2022

Joseph Kezele, M.D., will gave a presentation titled,
"WHERE DID THOSE THOUSANDS OF LANGUAGES COME FROM?" - A review of the structures of the body and the software needed for language are presented. This is followed by an outline of the evolutionary model of language development, the structure and shift of language sounds, the rules of language known as grammar, semantics, syntax and other factors. Then a world tour of language maps and the Biblical account of the origin of languages are presented.(Check back later. We expect to be able to provide link to a video recording of this presentation in the near future.)
About our speaker: Joseph Kezele, MD, studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomate of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians. He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, overseeing six creation science chapters throughout Arizona. He is Associate Professor at Arizona Christian University in Phoenix, teaching Human Genetics, Cell and Molecular Biology, Pathophysiology, Organic Chemistry and Creation Science Apologetics. Dr, Kezele has also been named a Logos Research Associate. He serves as Chairman of Deacons and teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona. He has participated in over 21 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, and Armenia. Dr. Kezele is one of the principle presenters in the Institute for Creation Research’s 4 DVD series, " Made in His Image". Four years ago, he returned to the Ukraine for three intense weeks of teaching Creation Science Apologetics and Evangelism as part of the activities prompted by the proclamation of the President of the Ukraine recognizing the 500th anniversary of Martin Luther's sparking the Reformation in Germany.
Dr. Kezele is a Logos Research Associate. He is also a Fellow of the Creation Research Society. Dr. Kezele is President of the Arizona Origins Science Association (AzOSA), and oversees the operations of 8 (soon to be 10) AzOSA chapters, one in each of 10 of Arizona's 15 counties.
"WHERE DID THOSE THOUSANDS OF LANGUAGES COME FROM?" - A review of the structures of the body and the software needed for language are presented. This is followed by an outline of the evolutionary model of language development, the structure and shift of language sounds, the rules of language known as grammar, semantics, syntax and other factors. Then a world tour of language maps and the Biblical account of the origin of languages are presented.(Check back later. We expect to be able to provide link to a video recording of this presentation in the near future.)
About our speaker: Joseph Kezele, MD, studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomate of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians. He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, overseeing six creation science chapters throughout Arizona. He is Associate Professor at Arizona Christian University in Phoenix, teaching Human Genetics, Cell and Molecular Biology, Pathophysiology, Organic Chemistry and Creation Science Apologetics. Dr, Kezele has also been named a Logos Research Associate. He serves as Chairman of Deacons and teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona. He has participated in over 21 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, and Armenia. Dr. Kezele is one of the principle presenters in the Institute for Creation Research’s 4 DVD series, " Made in His Image". Four years ago, he returned to the Ukraine for three intense weeks of teaching Creation Science Apologetics and Evangelism as part of the activities prompted by the proclamation of the President of the Ukraine recognizing the 500th anniversary of Martin Luther's sparking the Reformation in Germany.
Dr. Kezele is a Logos Research Associate. He is also a Fellow of the Creation Research Society. Dr. Kezele is President of the Arizona Origins Science Association (AzOSA), and oversees the operations of 8 (soon to be 10) AzOSA chapters, one in each of 10 of Arizona's 15 counties.
august 20, 2022

During our August 6 meeting, Dr. John Baumgardner demonstrated that the thesis of Materialism - that particles in motion and the forces between them are all that exist - is falsified by even a single example of something that is non-material, yet clearly exists. The example Dr. Baumgardner highlighted is language.
Mathematics has been called, "The Language of the Universe."
Our August 20 program, the PBS Nova documentary,
"THE GREAT MATH MYSTERY" allows us to watch evolutionary secular scientists wrestle with cognitive dissonance in their naturalistic worldviews as they struggle to answer the question, Is math discovered or invented? Along the way, the documentary provides many mind-boggling incidences of discoveries in abstract mathematics, which led to predictions about the material universe - predictions that were eventually confirmed by scientific observation.
Weren't able to attend, or want to see it again? This NOVA program is available on YouTube at https://www.youtube.com/watch?v=8mve0UoSxTo&list=PLRxEhF7If641_4Slm_A30DK8STmSnle01&index=1www.youtube.com/watch?v=8mve0UoSxTo&list=PLRxEhF7If641_4Slm_A30DK8STmSnle01&index=1
Mathematics has been called, "The Language of the Universe."
Our August 20 program, the PBS Nova documentary,
"THE GREAT MATH MYSTERY" allows us to watch evolutionary secular scientists wrestle with cognitive dissonance in their naturalistic worldviews as they struggle to answer the question, Is math discovered or invented? Along the way, the documentary provides many mind-boggling incidences of discoveries in abstract mathematics, which led to predictions about the material universe - predictions that were eventually confirmed by scientific observation.
Weren't able to attend, or want to see it again? This NOVA program is available on YouTube at https://www.youtube.com/watch?v=8mve0UoSxTo&list=PLRxEhF7If641_4Slm_A30DK8STmSnle01&index=1www.youtube.com/watch?v=8mve0UoSxTo&list=PLRxEhF7If641_4Slm_A30DK8STmSnle01&index=1
AUGUST 6, 2022

John Baumgardner, Ph.D., Geophysics and Space Physics, will gave a remote presentation by ZOOM titled, A LINGUISTIC ARGUMENT FOR GOD’S EXISTENCE. <<<< CLICK here for video - (Please be patient. Large file may take a minute to load. It is worth the wait!)
(This lecture was based on a 2015 paper by the same name found here - https://www.etsjets.org/files/JETS-PDFs/58/58-4/JETS_58-4_771-86_Baumgardner&Lyon.pdf )
Philosophical materialism is predicated on the claim that no non-material realities exist. Hence, a single substantive counterexample of this foundational truth claim falsifies that framework.
A category of reality which plays a huge role in the world around us is that of formal language. Formal language involves the assignment of snippets of meaning to a set of arbitrary symbols to form a vocabulary. It further involves a set of rules by which elements from the vocabulary may be joined together to form more complex meaning structures. Formal language includes not only human languages but also machine languages, mathematics, and the genetic specifications found in the DNA of living organisms.
Formal language in its ultimate essence is nothing more than encoded meaning. Because meaning is abstract and non-material, so is formal language in all its diverse expressions. Formal language plays such prominent roles in the world around us that it is impossible to argue that it is not real. Living systems are unthinkable apart from the non-material linguistic specifications on which they rely. Our thoughts and communications with others are also manifestations of non-material formal language but nonetheless are indisputably real. Thus, the claim that non-material realities do not exist is easy to demonstrate to be false. Because matter shows no innate ability to assign meaning to symbols or to manipulate such symbols, linguistic entities require a non-material source. The only candidate source is mind. The size and complexity of the linguistic structures on which living organisms rely demand a mind
for which there is but one viable candidate, the God of the Bible.
Reference: Baumgardner, J.R. and J.D. Lyon. 2015. A linguistic argument for God’s existence. JETS 58/4
771–86. https://answersingenesis.org/is-god-real/linguistic-argument-gods-existence/
About our speaker:
Education
Ph.D. Geophysics & Space Physics, 1983 - University of California, Los Angeles
M.S. Geophysics & Space Physics, 1981 - University of California, Los Angeles
M.S. Electrical Engineering, 1970 - Princeton University
B.S. Electrical Engineering, 1968 - Texas Tech University
Professional Positions
Liberty University, Research Professor Emeritus, School of Engineering, 4/2019-present
Logos Research Associates, Senior Research Associate, 10/2008-present. President 10/2008-1/2010, Vice President 1/2010-1/2022, Director at large, 1/2022-present.
Southern California Seminary, El Cajon, California, Adjunct Faculty, 3/2014-4/2018.
Department of Earth and Environmental Sciences, Ludwig Maximilian University, Munich, Germany, Adjunct Research Scientist, 01/2011 to 01/2014.
Institute for Creation Research, Associate Professor of Geophysics, 1/05-6/08.
Los Alamos National Laboratory, Theoretical Division, Technical Staff Member, 2/84 to 6/04.
Rockwell International, Rocketdyne Division, Laser Dept., Member of Technical Staff, 6/78-1/84.
Campus Crusade for Christ, Campus Ministry Staff, 7/75-6/78.
U. S. Air Force, Air Force Weapons Lab, Laser Division, Kirtland AFB, NM, Project Officer, 1/71-5/75.
Brief Personal History
After graduating from high school, I obtained a bachelor’s degree in electrical engineering from Texas Tech and then a master’s degree in electrical engineering from Princeton. But the most significant happening in my life before or since was my conversion to Christ through a verse-by-verse study of the gospel of John in a college Sunday school class in 1970. I liken this event to a curtain being pulled back on a dimension of reality I had earlier not even suspected to exist. Christ has since made my life an incredibly exciting and fulfilling experience—way beyond anything I otherwise might have imagined.
Soon after my conversion I began a four-year tour of active duty in the U. S. Air Force, a time of exciting and deep spiritual growth. A desire for Christian ministry then led me to join the campus ministry of Campus Crusade for Christ. Grieved by the deliberate use of evolution to assault and destroy the faith of Christian college students that I observed on the campus where I served, I began to present classroom lectures and evening forums to expose evolution’s fraudulent claims. In researching issues relating to the Genesis Flood and earth history in preparation for some of these events, I realized that Noah’s Flood logically involved planetary scale tectonic catastrophe. This awareness prompted me to leave Campus Crusade in 1978 to begin a Ph.D. program in geophysics at UCLA to obtain the training and credentials specifically to address the problem of the mechanism of the Genesis Flood at a professional scientific level. My Ph.D. thesis research involved the development of a 3-D spherical finite element model for the earth’s mantle, a program now known as TERRA. Most of this thesis research I conducted at Los Alamos National Laboratory. I completed my Ph.D. from UCLA in geophysics and space physics in 1983.
Between 1984 and 2004 I served as a staff scientist in the Theoretical Division at Los Alamos where, in addition to other computational fluid dynamics research, I was able to continue my research in planetary mantle dynamics, including the potential for catastrophic mantle overturn. I have presented my research results on catastrophic plate tectonics as the mechanism for Noah’s Flood at seven International Conferences on Creationism held in Pittsburgh beginning in 1986. Between 1997 and 2005 I served as a member of the Radioisotopes and the Age of the Earth (RATE) team that uncovered strong radioisotope evidence that the earth is thousands, not billions, of years old. One such line of evidence is the retention in zircon crystals in granite of more than a billion years’ worth, at present rates of decay, of radiogenic helium. Diffusion limits the lifetime of helium in these zircon crystals to only a few thousand years. Another line of evidence for a youthful earth was our discovery of C-14 in natural diamond.
Between 2005 and 2008 I served as a full-time staff scientist at the Institute for Creation Research in Santee, California. Much of my work there involved, as part of a small team, the development of Mendel’s Accountant, a state-of-the-art population genetics program for studying issues of mutation and selection in natural populations. This software shows, with no room for uncertainty, that natural selection does not and cannot deliver the miraculous results that evolutionists routinely attribute to it. Since 2008 I have been a senior research associate with Logos Research Associates (http://www.logosresearchassociates.org) where I continue to research topics relating to earth history and Scripture. In addition, from 2014 to 2018 I served as an adjunct faculty member at Southern California Seminary located in El Cajon, California, in its Center for Creation Studies. I currently serve as research professor emeritus in the School of Engineering at Liberty University. My wife Mary and I reside near Lynchburg, Virginia.
Highlights of My Technical Career
• As a project officer during four years of active duty at the Air Force Weapons Laboratory, Kirtland AFB, NM, I had responsibilities in the design of the resonator optics for a large, classified CO2 gas dynamics laser that was later built and flown in the Airborne Laser Laboratory, a refitted KC-135 aircraft. As part of this effort I developed a 3D optics/power extraction code that solved the gas kinetics rate equations in the supersonic flow where the lasing and power extraction occurred and computed the optical quality of output beam as affected by shocks in the supersonic flow. The laser performed according to specs and successfully demonstrated the capability from an airborne platform to destroy incoming missiles.
• As part of my Ph.D. thesis research at UCLA I developed the 3-D spherical code now known as TERRA for modeling the dynamics of planetary mantles. This code utilized the then newly discovered multigrid method for solving the large elliptic systems of equations arising in this application. The efficiency of multigrid plus the advantages of the almost uniform icosahedral grid made such 3-D calculations feasible for the first time. This code was the first 3-D spherical code of its kind and continues almost 30 years later to be the state of the art and used by several research groups around the world. One example is the research collaboration with Prof. Uwe Walzer and his geodynamics group at Friedrich Schiller University in Jena, Germany. The many diverse applications of TERRA by this group and the resulting publications may be found at www.geodyn.uni-jena.de. Another example of TERRA’s ongoing application is the research by Prof. Hans-Peter Bunge and his geodynamics group at Ludwig Maximilians University in Munich, Germany. Their publications can be found at https://www.geophysik.uni-muenchen.de/Members/bunge.
• During my 20 years in a computational fluid dynamics research group at Los Alamos National Laboratory, I was one of the main developers of the Pagosa code, a massively parallel multi-material Eulerian code that tracked material interfaces cell by cell in a robust high-order manner. The unique ability of this code at that time to treat a DOE mission-critical weapons application earned our team a Distinguished Performance Award in 1991.
• While at Los Alamos, I assisted the German Weather Service 1995-1999 in applying icosahedral grid techniques for modeling fluid flow on the sphere to the task of developing their next-generation weather forecast model named GME, a model that became operational in 2000 and is currently used in more than 20 countries.
• In 1996 I was part of a team of earth scientists who wrote a successful proposal for NASA’s High Performance Computing and Communication Initiative Round 2. This $1.3M award supported TERRA research from 1997-2000 and significantly advanced TERRA’s abilities. TERRA readily achieved NASA’s Round 2 massively parallel computing benchmarks.
• Also at Los Alamos, I was part of the ocean modeling team that developed the POP global ocean model, which is the ocean component of the NCAR Community Climate System Model, one of the two main U.S. climate models.
• During my tenure at Los Alamos, I supervised several graduate students who utilized my Terra mantle dynamics code for their Ph.D. thesis research.
• After retirement from Los Alamos, I have collaborated with Dr. John Sanford, a Cornell geneticist, to develop a state-of-the-art population genetics program named Mendel’s Accountant. I have also advised additional graduate students who have utilized the Terra software in their Ph.D. thesis research. In addition I have undertaken research into the manner that rapid plate tectonics during the Genesis Flood likely generated tens of thousands of giant tsunamis that played a key role in the erosion, transport, and deposition of the fossil-bearing sediment layers that blanket the continents today.
• Over 80 peer-reviewed publications.
Selected Publications
H. E. Cho, Y. Hammi, A. L. Bowman, S. Karato, J. R. Baumgardner, M. F. Horstemeyer, “A unified static and dynamic recrystallization Internal State Variable (ISV) constitutive model coupled with grain size evolution for metals and mineral aggregates,” International Journal of Plasticity, 112, pp. 123-157, DOI: 10.1016/j.ijplas.2018.08.009, 2019.
https://www.sciencedirect.com/science/article/pii/S0749641918303139
J. Baumgardner, “Understanding how the Flood sediment record was formed: The role of large tsunamis,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 287–305, Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1020&context=icc_proceedings
N. Cho, J. Baumgardner, J.A. Sherburn, and M.F. Horstemeyer, “Numerical investigation of strength-reducing mechanisms of mantle rock during the Genesis Flood,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 707–730, Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1067&context=icc_proceedings
T. G. Tenev, J. Baumgardner, and M. F. Horstemeyer, “A solution for the distant starlight problem using creation time coordinates,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 82-94. Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1017&context=icc_proceedings
J. Sanford, R. Carter, W. Brewer, J. Baumgardner, B. Potter, and J. Potter, “Adam and Eve, designed diversity, and allele frequencies,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 200–216. Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1079&context=icc_proceedings
J. Baumgardner, “Numerical modeling of the large-scale erosion, sediment transport, and deposition processes of the Genesis Flood,” Answers Research Journal 11:149–170, 2018
https://assets.answersingenesis.org/doc/articles/pdf-versions/arj/v11/numerical-modeling-sediment-transport.pdf
J. R. Baumgardner and J. D. Lyon, “A linguistic argument for God’s existence,” Journal of the Evangelical Theological Society 58/4 771–786, 2015.
http://media.wix.com/ugd/a704d4_87dbd5d9be6944618e01950af43559e3.pdf
J. Sanford, W. Brewer, F. Smith and J. Baumgardner, “The waiting time problem in a model hominin population,” Theoretical Biology and Medical Modelling 12:18, 2015.
https://tbiomed.biomedcentral.com/articles/10.1186/s12976-015-0016-z
J. A. Sherburn, J. R. Baumgardner, and M. F. Horstemeyer, “New material model reveals inherent tendency in mantle minerals for runaway mantle dynamics,” in Proceedings of the Seventh International Conference on Creationism, M. Horstemeyer, editor, Creation Science Fellowship, Inc., Pittsburgh, PA, 2013.
http://media.wix.com/ugd/a704d4_131cfb5ab517b7c4f5c9ee0fab5ca351.pdf
J. R. Baumgardner, W. H. Brewer, and J. C. Sanford, “Can synergistic epistasis halt mutation accumulation? Results from numerical simulation,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 312-337, 2013.
http://marksmannet.com/RobertMarks/REPRINTS/BINP/9789814508728_0013.pdf
W. H. Brewer, J. R. Baumgardner, and J. C. Sanford, “Using numerical simulation to test the ‘mutation-count’ hypothesis,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 298-311, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0012.pdf
J. C. Sanford, J.R. Baumgardner, W. H. Brewer, “Selection threshold severely constrains capture of beneficial mutations,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 264-297, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0011.pdf
P. Gibson, J. R. Baumgardner, W. H. Brewer, and J. C. Sanford, “Can purifying natural selection preserve biological information?” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J. C. Sanford, editors, World Scientific, Singapore, 232-263, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0010.pdf
J. Baumgardner, “Global tectonics—clarity, not confusion,” Journal of Creation 27(1), 99-106, 2013.
http://creation.com/images/pdfs/tj/j27_1/j27_1_99-106.pdf
J. Baumgardner, “Do radioisotope methods yield trustworthy relative ages for the earth’s rocks?” Journal of Creation 26(3), 68-75, 2012.
http://creation.com/images/pdfs/tj/j26_3/j26_3_68-75.pdf
J. Baumgardner, “Is plate tectonics occurring today?” Journal of Creation 26(1), 101-105, 2012.
http://creation.com/plate-tectonics-today
J. Baumgardner, “Language, complexity, and design,” in Divine Action and Natural Selection: Science, Faith and Evolution, J. Seckback and R. Gordon, eds., World Scientific, Singapore, 939-980, 2009.
http://media.wix.com/ugd/a704d4_a84beb1f9e4aefec9f4f9444eeec8550.pdf
J. Baumgardner, J. Sanford, W. Brewer, P. Gibson, and W. ReMine, “Mendel’s Accountant: A new population genetics simulation tool for studying mutation and natural selection,” in Proceedings of the Sixth International Conference on Creationism, A. A. Snelling, editor, Creation Science Fellowship, Pittsburgh, PA, 87-98, 2008.
http://www.icr.org/i/pdf/technical/Mendels-Accountant.pdf
J. Sanford, J. Baumgardner, J. Sanford, W. Brewer, W. ReMine, and P. Gibson, “Using numerical simulation to test the validity of neo-Darwinian theory,” in Proceedings of the Sixth International Conference on Creationism, A. A. Snelling, editor, Creation Science Fellowship, Pittsburgh, PA, 165-176, 2008.
http://www.icr.org/i/pdf/technical/Using-Numerical-Simulation-to-Test-the-Validity-of-Neo-Darwinian-Theory.pdf
J. Sanford, J. Baumgardner, W. Brewer, P. Gibson, and W. ReMine, “Mendel's Accountant: a biologically realistic forward-time population genetics program,” Scalable Computing: Practice and Experience, 8(2), 147-165, 2007.
https://www.researchgate.net/publication/220101430_Mendel's_Accountant_A_biologically_realistic_forward-time_population_genetics_program
U. Walzer, R. Hendel, and J. Baumgardner, “Whole-mantle convection, continent generation, and preservation of geochemical heterogeneity,” in High Performance Computing in Science and Engineering ’07, W. Nagel, D. Kröner, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, IBSN 978-3-540-74738-3, 603-645, 2007.
https://www.researchgate.net/publication/227165432_Whole-Mantle_Convection_Continent_Generation_and_Preservation_of_Geochemical_Heterogeneity
J. Sanford, J. Baumgardner, W. Brewer, P. Gibson, and W. ReMine, “Using computer simulation to understand mutation accumulation dynamics and genetic load,” in Y. Shi et al. (eds.), ICCS 2007, Part II, Lecture Notes in Computer Science, 4488, Springer-Verlag, Berlin, Heidelberg, pp. 386-392, 2007.
https://www.researchgate.net/publication/220859353_Using_Computer_Simulation_to_Understand_Mutation_Accumulation_Dynamics_and_Genetic_Load
U. Walzer, R. Hendel, and J. Baumgardner, “Continental growth and thermal convection in the Earth’s mantle,” in High Performance Computing in Science and Engineering ’06, W. Nagel, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 978-3-540-36165-7, 473-497, 2006
K.-D. Gottschaldt, U. Walzer, R. F. Hendel, D. R. Stegman, J. R. Baumgardner, and H.-B. Muhlhaus, “Stirring in 3-d spherical models of convection in the Earth's mantle,” Philosophical Magazine, 86, 3175-3204, 2006.
J. R. Baumgardner, “14C Evidence for a Recent Global Flood and a Young Earth,” in Radioisotopes and the Age of the Earth: Results of a Young-Earth Creationist Research Initiative, Volume II, L. Vardiman, A. Snelling, and E. Chaffin, eds., Institute for Creation Research, Santee, CA, 587-630, 2005.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Scaling laws for time-dependent stagnant lid convection in a spherical shell,” Phys. Earth Planet. Inter., 149, 361-370, 2005.
U. Walzer, R. Hendel, and J. Baumgardner, “Plateness of the oceanic lithosphere and the thermal evolution of the Earth's mantle,” in High Performance Computing in Science and Engineering '05, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3540283773, 289-304, 2005.
U. Walzer, R. Hendel, and J. Baumgardner, “The effects of a variation of the radial viscosity profile on mantle evolution,” Tectonophysics, 384, 55-90, 2004.
C. C. Reese, V. S. Solomatov, J. R. Baumgardner, D. R. Stegman, and A. V. Vezolainen, “Magmatic evolution of impact-induced Martian mantle plumes and the origin of Tharsis,” J. Geophys. Res., 109, 10.1029/2003 JE002222, 2004.
U. Walzer, R. Hendel, and J. Baumgardner, “Toward a geochemical model for the evolution of the Earth’s mantle,” in High Performance Computing in Science and Engineering '04, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-22943-4, 395-454, 2004.
U. Walzer, R. Hendel, and J. Baumgardner, “Generation of plate-tectonic behavior and a new viscosity profile of the Earth's mantle,” in NIC Symposium 2004 Proceedings, D. Wolf, G. Münster, and M. Kremer (eds.), NIC Series 20, 419-428., ISBN 3-00-012372-5, 2003.
J. R. Baumgardner, “Catastrophic plate tectonics: the physics behind the Genesis Flood,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., Editor, Creation Science Fellowship, Pittsburgh, PA, 113-126, 2003.
D. R. Humphreys, S. A. Austin, J. R. Baumgardner, and A. A Snelling, “Helium diffusion rates support accelerated nuclear decay,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 175-196, 2003.
L. Vardiman, S. A. Austin, J. R. Baumgardner, E. F. Chafin, D. B. DeYoung, D. R. Humphreys and A. A Snelling, “Radioisotopes and the age of the Earth,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 337-348, 2003.
U. Walzer, R. Hendel, and J. Baumgardner, “Viscosity stratification and a 3-D compressible spherical shell model of mantle evolution”, in High Performance Computing in Science and Engineering '03, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-40850-9, 27-67, 2003.
D. R. Stegman, A.M. Jellinek, S. A. Zatman, J. R. Baumgardner, and M. A. Richards, “An early lunar core dynamo driven by thermochemical mantle convection,” Nature, 421, 143-146, 2003.
M. F. Horstemeyer and J. R. Baumgardner, “What initiated the Flood cataclysm?” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 155-164, 2003.
J. R. Baumgardner, “Catastrophic plate tectonics: the geophysical context of the Genesis Flood,” “Dealing carefully with the data,” and “A constructive quest for truth,” all contributions to a “Forum on Catastrophic Plate Tectonics,” Ex Nihilo Technical Journal, 16, 57-85, 2002.
(http://www.answersingenesis.org/home/area/magazines/tj/docs/16n1p57Forum.asp)
H.-P. Bunge, M. A. Richards, and J. R. Baumgardner, “Mantle-circulation models with sequential data assimilation: inferring present-day mantle structure from plate-motion histories,” Phil. Trans. R. Soc. Lond. A., 360, 2545-2567, 2002.
D. A. Randall, T. D. Ringler, R. P. Heikes, P. Jones, and J. Baumgardner, “Climate Modeling with Spherical Geodesic Grids,” Computing in Science and Engineering, 4(5), 32-41, 2002.
U. Walzer, R. Hendel, and J. Baumgardner, “Variation of non-dimensional numbers and a thermal evolution model of the Earth's mantle,” in High Performance Computing in Science and Engineering '02, E. Krause and W. Jäger, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-43860-2, 89-103, 2002.
D. R. Stegman, M.A. Richards, and J.R. Baumgardner, Effects of depth dependent viscosity and plate motions on maintaining a relatively uniform MORB reservoir in whole mantle flow, J. Geophys. Res., 107, (B6), 2116, doi:10.1029/2001JB000192, 2002.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Survival of impact-induced thermal anomalies in the Martian mantle,” J. Geophys. Res.- Planets, 107(10), 5082-5092, 2002.
D. Majewski, D. Liermann, P. Prohl, B. Ritter, M. Buchhold, T. Hanisch, G. Paul, W. Wergen, and J. Baumgardner, “The global icosahedral-hexagonal grid point model GME: Operational version and high resolution tests,” Mon. Wea. Rev., 130, 319-338, 2002.
M. A. Richards, W.-S. Yang, and J. R. Baumgardner, “The role of a low viscosity zone in stabilizing plate tectonics: implications for comparative terrestrial planetology,” Geochem., Geophys., Geosys., 2, 2001.
J. R. Baumgardner, “Distribution of Radioactive Isotopes in the Earth” in Radioisotopes and the Age of the Earth: A Young Earth Creation Research Initiative, Larry Vardiman, Andrew A. Snelling, and Eugene F. Chaffin, eds., Institute for Creation Research, Santee, CA, 49-94, 2000.
J. K. Dukowicz and J. R. Baumgardner, “Incremental remapping as a transport/advection algorithm,” J. Comp. Phys., 160, 318-335, 2000.
W.-S. Yang and J. R. Baumgardner, "Matrix-dependent transfer multigrid method for strongly variable viscosity infinite Prandtl number thermal convection," Geophys. and Astrophys. Fluid Dyn., 92, 151-195, 2000.
H. R. Wenk, J. R. Baumgardner, C. N. Tome, and R. Lebensohn, "A deformation model to explain anisotropy of the inner core," J. Geophys. Res., 105, 5663-5677, 2000.
M. A. Richards, H.-P. Bunge, C. Lithgow-Bertelloni, and J. R. Baumgardner, "Mantle convection and plate motion history: Toward general circulation models," History and Dynamics of Global Plate Motions, AGU Monograph Series, 1999.
M. A. Richards, H.-P. Bunge, Y. Ricard, and J. R. Baumgardner, "Polar wandering in mantle convection models," Geophys. Res. Lett., 26, 1777-1780, 1999.
H.-P. Bunge, M. A. Richards, C. Lithgow-Bertelloni, J. R. Baumgardner, S. P. Grand, and B. A. Romanowicz, "Time scales and heterogeneity structure in geodynamic earth models," Science, 280, 91-95, 1998.
Hans-Peter Bunge, Mark A. Richards, and John R. Baumgardner, "A sensitivity study of 3-D spherical mantle convection at 108 Rayleigh number: effects of depth-dependent viscosity, heating mode, and an endothermic phase change," J. Geophys. Res.,102, B6, 11991-12007, 1997.
Hans-Peter Bunge, Mark A. Richards, and John R. Baumgardner, "The effect of depth-dependent viscosity on the planform of mantle convection," Nature, 379, 436-438, 1996.
Hans-Peter Bunge and John R. Baumgardner, "Mantle convection modeling on parallel virtual machines," Computers in Physics, 9, 207-215, 1995.
S. A. Austin, J. R. Baumgardner, D. R. Humphreys, A. A. Snelling, L. Vardiman, and K. P. Wise, "Catastrophic Plate Tectonics: A Global Flood Model of Earth History," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 609-621, 1994.
J. R. Baumgardner, "Computer Modeling of the Large-Scale Tectonics Associated with the Genesis Flood," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 49-62, 1994.
J. R. Baumgardner, "Runaway Subduction as the Driving Mechanism for the Genesis Flood," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 63-75, 1994.
D. B. Kothe, J. R. Baumgardner, J. H. Cerutti, B. J. Daly, K. S. Holian, E. M. Kober, S. J. Mosso, J. W. Painter, R. D. Smith, and M. D. Torrey, "PAGOSA: A massively-parallel, multi-material hydrodynamics model for three-dimensional high-speed flow and high-rate material deformation," Proceedings of the 1993 Conference on High Performance Computing: Grand Challenges in Computer Simulation, Society for Computer Simulation, 9-14, 1993.
John R. Baumgardner, "3-D numerical investigation of the mantle dynamics associated with the breakup of Pangea," in Flow and Creep in the Solar System: Observations, Modeling, and Theory, D. B. Stone and S. K. Runcorn, eds., NATO ASI Series C, Vol. 391, 207-224, 1993.
M. A. Moreno, G. Schubert, J. Baumgardner, M. G. Kivelson, and D. A. Paige, "Io's volcanic and sublimation atmospheres," Icarus, 93, 63-81, 1991.
John R. Baumgardner, "Application of supercomputers to 3-D mantle convection," in The Physics of the Planets, S. K. Runcorn, ed., John Wiley and Sons, 199-231, 1988.
J. Baumgardner, M. A. Moreno, G. Schubert, and M. G. Kivelson, "Two classes of volcanic eruptions and their corresponding atmospheres on Io," Bull. Am. Astr. Assoc., 19(3), 856, 1987.
John R. Baumgardner, "Three-dimensional treatment of convective flow in the earth's mantle," J. Stat. Phys., 39, 501-511, 1985.
John R. Baumgardner and Paul O. Frederickson, "Icosahedral discretization of the two-sphere," SIAM J. Numer. Anal., 22, 1107-1115, 1985.
Selected Conference Abstracts
D. R. Stegman, M. Jellinek, J. Baumgardner, and M. Richards, “A mantle dynamics perspective on generating an early lunar dynamo,” (abstract) Eos, Trans. Am. Geophys. Union, 84, (2003 Fall Meeting Supplement), 2003.
D. R. Humphreys, J. R. Baumgardner, A. A. Snelling, and S. A. Austin, “Recently measured helium diffusion rate for zircon suggests inconsistency with U-Pb age for Fenton Hill granodiorite,” (abstract) Eos, Trans. Am. Geophys. Union, 84, (2003 Fall Meeting Supplement), 2003.
J. R. Baumgardner, Y. Yamagishi, and D.R. Stegman, “Efficient visual data exploration in high-resolution global dynamical models of the Earth's mantle,” (abstract) Eos, Trans. Am. Geophys. Union, 83, (2002 Fall Meeting Supplement), 2002.
J. R. Baumgardner, “3D numerical modeling of the dynamics of Earth's mantle,” Am. Phys. Soc. Four Corners Annual Meeting, Las Cruces, New Mexico, 2001.
J. R. Baumgardner, W.-S. Yang, and M. A. Richards, “The role of a low viscosity zone in stabilizing a plate tectonics style of mantle convection,” (abstract) Eos, Trans. Am. Geophys. Union, 81, (2000 Fall Meeting Supplement), 2000.
J. R. Baumgardner and W.-S. Yang, “Earthlike mantle convection from relatively simple rheology,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F26, 1999.
M. A. Richards, W.-S. Yang, and J. R. Baumgardner, “The effectiveness of finite yield stress in obtaining platelike surface velocities,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F962, 1999.
D. R. Stegman, M. A. Richards, and J. R. Baumgardner, “A parallel implementation of Lagrangian tracers in TERRA,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F950, 1999.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Impacts and the thermal evolution of Mars,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F618, 1999.
John R. Baumgardner, Mark A. Richards, Woo-Sun Yang, and Carolina R. Lithgow-Bertelloni, "3-D Spherical Models of Plate Motion With Laterally Varying Rheology," (abstract) Eos, Trans. Am. Geophys. Union, 79, (1998 Fall Meeting Supplement), F911, 1998.
John R. Baumgardner and Woo-Sun Yang, "A finite element multigrid formulation for variable viscosity in 3-D spherical geometry," (abstract) Eos, Trans. Am. Geophys. Union, 77, (Fall Meeting Supplement), F750, 1996.
J. R. Baumgardner, "Thermal runaway in the mantle" (abstract) Eos, Trans. Am. Geophys. Union, 75, 687, 1994.
John Baumgardner, "3-D numerical investigation of the mantle dynamics associated with the breakup of Pangea," (abstract) Eos, Trans. Am. Geophys. Union, 73, 1992 Fall Meeting Abstract Volume, 576-577, 1992.
(This lecture was based on a 2015 paper by the same name found here - https://www.etsjets.org/files/JETS-PDFs/58/58-4/JETS_58-4_771-86_Baumgardner&Lyon.pdf )
Philosophical materialism is predicated on the claim that no non-material realities exist. Hence, a single substantive counterexample of this foundational truth claim falsifies that framework.
A category of reality which plays a huge role in the world around us is that of formal language. Formal language involves the assignment of snippets of meaning to a set of arbitrary symbols to form a vocabulary. It further involves a set of rules by which elements from the vocabulary may be joined together to form more complex meaning structures. Formal language includes not only human languages but also machine languages, mathematics, and the genetic specifications found in the DNA of living organisms.
Formal language in its ultimate essence is nothing more than encoded meaning. Because meaning is abstract and non-material, so is formal language in all its diverse expressions. Formal language plays such prominent roles in the world around us that it is impossible to argue that it is not real. Living systems are unthinkable apart from the non-material linguistic specifications on which they rely. Our thoughts and communications with others are also manifestations of non-material formal language but nonetheless are indisputably real. Thus, the claim that non-material realities do not exist is easy to demonstrate to be false. Because matter shows no innate ability to assign meaning to symbols or to manipulate such symbols, linguistic entities require a non-material source. The only candidate source is mind. The size and complexity of the linguistic structures on which living organisms rely demand a mind
for which there is but one viable candidate, the God of the Bible.
Reference: Baumgardner, J.R. and J.D. Lyon. 2015. A linguistic argument for God’s existence. JETS 58/4
771–86. https://answersingenesis.org/is-god-real/linguistic-argument-gods-existence/
About our speaker:
Education
Ph.D. Geophysics & Space Physics, 1983 - University of California, Los Angeles
M.S. Geophysics & Space Physics, 1981 - University of California, Los Angeles
M.S. Electrical Engineering, 1970 - Princeton University
B.S. Electrical Engineering, 1968 - Texas Tech University
Professional Positions
Liberty University, Research Professor Emeritus, School of Engineering, 4/2019-present
Logos Research Associates, Senior Research Associate, 10/2008-present. President 10/2008-1/2010, Vice President 1/2010-1/2022, Director at large, 1/2022-present.
Southern California Seminary, El Cajon, California, Adjunct Faculty, 3/2014-4/2018.
Department of Earth and Environmental Sciences, Ludwig Maximilian University, Munich, Germany, Adjunct Research Scientist, 01/2011 to 01/2014.
Institute for Creation Research, Associate Professor of Geophysics, 1/05-6/08.
Los Alamos National Laboratory, Theoretical Division, Technical Staff Member, 2/84 to 6/04.
Rockwell International, Rocketdyne Division, Laser Dept., Member of Technical Staff, 6/78-1/84.
Campus Crusade for Christ, Campus Ministry Staff, 7/75-6/78.
U. S. Air Force, Air Force Weapons Lab, Laser Division, Kirtland AFB, NM, Project Officer, 1/71-5/75.
Brief Personal History
After graduating from high school, I obtained a bachelor’s degree in electrical engineering from Texas Tech and then a master’s degree in electrical engineering from Princeton. But the most significant happening in my life before or since was my conversion to Christ through a verse-by-verse study of the gospel of John in a college Sunday school class in 1970. I liken this event to a curtain being pulled back on a dimension of reality I had earlier not even suspected to exist. Christ has since made my life an incredibly exciting and fulfilling experience—way beyond anything I otherwise might have imagined.
Soon after my conversion I began a four-year tour of active duty in the U. S. Air Force, a time of exciting and deep spiritual growth. A desire for Christian ministry then led me to join the campus ministry of Campus Crusade for Christ. Grieved by the deliberate use of evolution to assault and destroy the faith of Christian college students that I observed on the campus where I served, I began to present classroom lectures and evening forums to expose evolution’s fraudulent claims. In researching issues relating to the Genesis Flood and earth history in preparation for some of these events, I realized that Noah’s Flood logically involved planetary scale tectonic catastrophe. This awareness prompted me to leave Campus Crusade in 1978 to begin a Ph.D. program in geophysics at UCLA to obtain the training and credentials specifically to address the problem of the mechanism of the Genesis Flood at a professional scientific level. My Ph.D. thesis research involved the development of a 3-D spherical finite element model for the earth’s mantle, a program now known as TERRA. Most of this thesis research I conducted at Los Alamos National Laboratory. I completed my Ph.D. from UCLA in geophysics and space physics in 1983.
Between 1984 and 2004 I served as a staff scientist in the Theoretical Division at Los Alamos where, in addition to other computational fluid dynamics research, I was able to continue my research in planetary mantle dynamics, including the potential for catastrophic mantle overturn. I have presented my research results on catastrophic plate tectonics as the mechanism for Noah’s Flood at seven International Conferences on Creationism held in Pittsburgh beginning in 1986. Between 1997 and 2005 I served as a member of the Radioisotopes and the Age of the Earth (RATE) team that uncovered strong radioisotope evidence that the earth is thousands, not billions, of years old. One such line of evidence is the retention in zircon crystals in granite of more than a billion years’ worth, at present rates of decay, of radiogenic helium. Diffusion limits the lifetime of helium in these zircon crystals to only a few thousand years. Another line of evidence for a youthful earth was our discovery of C-14 in natural diamond.
Between 2005 and 2008 I served as a full-time staff scientist at the Institute for Creation Research in Santee, California. Much of my work there involved, as part of a small team, the development of Mendel’s Accountant, a state-of-the-art population genetics program for studying issues of mutation and selection in natural populations. This software shows, with no room for uncertainty, that natural selection does not and cannot deliver the miraculous results that evolutionists routinely attribute to it. Since 2008 I have been a senior research associate with Logos Research Associates (http://www.logosresearchassociates.org) where I continue to research topics relating to earth history and Scripture. In addition, from 2014 to 2018 I served as an adjunct faculty member at Southern California Seminary located in El Cajon, California, in its Center for Creation Studies. I currently serve as research professor emeritus in the School of Engineering at Liberty University. My wife Mary and I reside near Lynchburg, Virginia.
Highlights of My Technical Career
• As a project officer during four years of active duty at the Air Force Weapons Laboratory, Kirtland AFB, NM, I had responsibilities in the design of the resonator optics for a large, classified CO2 gas dynamics laser that was later built and flown in the Airborne Laser Laboratory, a refitted KC-135 aircraft. As part of this effort I developed a 3D optics/power extraction code that solved the gas kinetics rate equations in the supersonic flow where the lasing and power extraction occurred and computed the optical quality of output beam as affected by shocks in the supersonic flow. The laser performed according to specs and successfully demonstrated the capability from an airborne platform to destroy incoming missiles.
• As part of my Ph.D. thesis research at UCLA I developed the 3-D spherical code now known as TERRA for modeling the dynamics of planetary mantles. This code utilized the then newly discovered multigrid method for solving the large elliptic systems of equations arising in this application. The efficiency of multigrid plus the advantages of the almost uniform icosahedral grid made such 3-D calculations feasible for the first time. This code was the first 3-D spherical code of its kind and continues almost 30 years later to be the state of the art and used by several research groups around the world. One example is the research collaboration with Prof. Uwe Walzer and his geodynamics group at Friedrich Schiller University in Jena, Germany. The many diverse applications of TERRA by this group and the resulting publications may be found at www.geodyn.uni-jena.de. Another example of TERRA’s ongoing application is the research by Prof. Hans-Peter Bunge and his geodynamics group at Ludwig Maximilians University in Munich, Germany. Their publications can be found at https://www.geophysik.uni-muenchen.de/Members/bunge.
• During my 20 years in a computational fluid dynamics research group at Los Alamos National Laboratory, I was one of the main developers of the Pagosa code, a massively parallel multi-material Eulerian code that tracked material interfaces cell by cell in a robust high-order manner. The unique ability of this code at that time to treat a DOE mission-critical weapons application earned our team a Distinguished Performance Award in 1991.
• While at Los Alamos, I assisted the German Weather Service 1995-1999 in applying icosahedral grid techniques for modeling fluid flow on the sphere to the task of developing their next-generation weather forecast model named GME, a model that became operational in 2000 and is currently used in more than 20 countries.
• In 1996 I was part of a team of earth scientists who wrote a successful proposal for NASA’s High Performance Computing and Communication Initiative Round 2. This $1.3M award supported TERRA research from 1997-2000 and significantly advanced TERRA’s abilities. TERRA readily achieved NASA’s Round 2 massively parallel computing benchmarks.
• Also at Los Alamos, I was part of the ocean modeling team that developed the POP global ocean model, which is the ocean component of the NCAR Community Climate System Model, one of the two main U.S. climate models.
• During my tenure at Los Alamos, I supervised several graduate students who utilized my Terra mantle dynamics code for their Ph.D. thesis research.
• After retirement from Los Alamos, I have collaborated with Dr. John Sanford, a Cornell geneticist, to develop a state-of-the-art population genetics program named Mendel’s Accountant. I have also advised additional graduate students who have utilized the Terra software in their Ph.D. thesis research. In addition I have undertaken research into the manner that rapid plate tectonics during the Genesis Flood likely generated tens of thousands of giant tsunamis that played a key role in the erosion, transport, and deposition of the fossil-bearing sediment layers that blanket the continents today.
• Over 80 peer-reviewed publications.
Selected Publications
H. E. Cho, Y. Hammi, A. L. Bowman, S. Karato, J. R. Baumgardner, M. F. Horstemeyer, “A unified static and dynamic recrystallization Internal State Variable (ISV) constitutive model coupled with grain size evolution for metals and mineral aggregates,” International Journal of Plasticity, 112, pp. 123-157, DOI: 10.1016/j.ijplas.2018.08.009, 2019.
https://www.sciencedirect.com/science/article/pii/S0749641918303139
J. Baumgardner, “Understanding how the Flood sediment record was formed: The role of large tsunamis,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 287–305, Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1020&context=icc_proceedings
N. Cho, J. Baumgardner, J.A. Sherburn, and M.F. Horstemeyer, “Numerical investigation of strength-reducing mechanisms of mantle rock during the Genesis Flood,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 707–730, Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1067&context=icc_proceedings
T. G. Tenev, J. Baumgardner, and M. F. Horstemeyer, “A solution for the distant starlight problem using creation time coordinates,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 82-94. Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1017&context=icc_proceedings
J. Sanford, R. Carter, W. Brewer, J. Baumgardner, B. Potter, and J. Potter, “Adam and Eve, designed diversity, and allele frequencies,” in Proceedings of the Eighth International Conference on Creationism, J. H. Whitmore, ed., pp. 200–216. Pittsburgh, Pennsylvania: Creation Science Fellowship, 2018.
https://digitalcommons.cedarville.edu/cgi/viewcontent.cgi?article=1079&context=icc_proceedings
J. Baumgardner, “Numerical modeling of the large-scale erosion, sediment transport, and deposition processes of the Genesis Flood,” Answers Research Journal 11:149–170, 2018
https://assets.answersingenesis.org/doc/articles/pdf-versions/arj/v11/numerical-modeling-sediment-transport.pdf
J. R. Baumgardner and J. D. Lyon, “A linguistic argument for God’s existence,” Journal of the Evangelical Theological Society 58/4 771–786, 2015.
http://media.wix.com/ugd/a704d4_87dbd5d9be6944618e01950af43559e3.pdf
J. Sanford, W. Brewer, F. Smith and J. Baumgardner, “The waiting time problem in a model hominin population,” Theoretical Biology and Medical Modelling 12:18, 2015.
https://tbiomed.biomedcentral.com/articles/10.1186/s12976-015-0016-z
J. A. Sherburn, J. R. Baumgardner, and M. F. Horstemeyer, “New material model reveals inherent tendency in mantle minerals for runaway mantle dynamics,” in Proceedings of the Seventh International Conference on Creationism, M. Horstemeyer, editor, Creation Science Fellowship, Inc., Pittsburgh, PA, 2013.
http://media.wix.com/ugd/a704d4_131cfb5ab517b7c4f5c9ee0fab5ca351.pdf
J. R. Baumgardner, W. H. Brewer, and J. C. Sanford, “Can synergistic epistasis halt mutation accumulation? Results from numerical simulation,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 312-337, 2013.
http://marksmannet.com/RobertMarks/REPRINTS/BINP/9789814508728_0013.pdf
W. H. Brewer, J. R. Baumgardner, and J. C. Sanford, “Using numerical simulation to test the ‘mutation-count’ hypothesis,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 298-311, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0012.pdf
J. C. Sanford, J.R. Baumgardner, W. H. Brewer, “Selection threshold severely constrains capture of beneficial mutations,” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J.C. Sanford, editors, World Scientific, Singapore, 264-297, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0011.pdf
P. Gibson, J. R. Baumgardner, W. H. Brewer, and J. C. Sanford, “Can purifying natural selection preserve biological information?” in Biological Information: New Perspectives, R. J. Marks II, M. J. Behe, W. A. Dembski, B. L. Gordon, and J. C. Sanford, editors, World Scientific, Singapore, 232-263, 2013.
http://robertmarks.org/REPRINTS/BINP/9789814508728_0010.pdf
J. Baumgardner, “Global tectonics—clarity, not confusion,” Journal of Creation 27(1), 99-106, 2013.
http://creation.com/images/pdfs/tj/j27_1/j27_1_99-106.pdf
J. Baumgardner, “Do radioisotope methods yield trustworthy relative ages for the earth’s rocks?” Journal of Creation 26(3), 68-75, 2012.
http://creation.com/images/pdfs/tj/j26_3/j26_3_68-75.pdf
J. Baumgardner, “Is plate tectonics occurring today?” Journal of Creation 26(1), 101-105, 2012.
http://creation.com/plate-tectonics-today
J. Baumgardner, “Language, complexity, and design,” in Divine Action and Natural Selection: Science, Faith and Evolution, J. Seckback and R. Gordon, eds., World Scientific, Singapore, 939-980, 2009.
http://media.wix.com/ugd/a704d4_a84beb1f9e4aefec9f4f9444eeec8550.pdf
J. Baumgardner, J. Sanford, W. Brewer, P. Gibson, and W. ReMine, “Mendel’s Accountant: A new population genetics simulation tool for studying mutation and natural selection,” in Proceedings of the Sixth International Conference on Creationism, A. A. Snelling, editor, Creation Science Fellowship, Pittsburgh, PA, 87-98, 2008.
http://www.icr.org/i/pdf/technical/Mendels-Accountant.pdf
J. Sanford, J. Baumgardner, J. Sanford, W. Brewer, W. ReMine, and P. Gibson, “Using numerical simulation to test the validity of neo-Darwinian theory,” in Proceedings of the Sixth International Conference on Creationism, A. A. Snelling, editor, Creation Science Fellowship, Pittsburgh, PA, 165-176, 2008.
http://www.icr.org/i/pdf/technical/Using-Numerical-Simulation-to-Test-the-Validity-of-Neo-Darwinian-Theory.pdf
J. Sanford, J. Baumgardner, W. Brewer, P. Gibson, and W. ReMine, “Mendel's Accountant: a biologically realistic forward-time population genetics program,” Scalable Computing: Practice and Experience, 8(2), 147-165, 2007.
https://www.researchgate.net/publication/220101430_Mendel's_Accountant_A_biologically_realistic_forward-time_population_genetics_program
U. Walzer, R. Hendel, and J. Baumgardner, “Whole-mantle convection, continent generation, and preservation of geochemical heterogeneity,” in High Performance Computing in Science and Engineering ’07, W. Nagel, D. Kröner, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, IBSN 978-3-540-74738-3, 603-645, 2007.
https://www.researchgate.net/publication/227165432_Whole-Mantle_Convection_Continent_Generation_and_Preservation_of_Geochemical_Heterogeneity
J. Sanford, J. Baumgardner, W. Brewer, P. Gibson, and W. ReMine, “Using computer simulation to understand mutation accumulation dynamics and genetic load,” in Y. Shi et al. (eds.), ICCS 2007, Part II, Lecture Notes in Computer Science, 4488, Springer-Verlag, Berlin, Heidelberg, pp. 386-392, 2007.
https://www.researchgate.net/publication/220859353_Using_Computer_Simulation_to_Understand_Mutation_Accumulation_Dynamics_and_Genetic_Load
U. Walzer, R. Hendel, and J. Baumgardner, “Continental growth and thermal convection in the Earth’s mantle,” in High Performance Computing in Science and Engineering ’06, W. Nagel, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 978-3-540-36165-7, 473-497, 2006
K.-D. Gottschaldt, U. Walzer, R. F. Hendel, D. R. Stegman, J. R. Baumgardner, and H.-B. Muhlhaus, “Stirring in 3-d spherical models of convection in the Earth's mantle,” Philosophical Magazine, 86, 3175-3204, 2006.
J. R. Baumgardner, “14C Evidence for a Recent Global Flood and a Young Earth,” in Radioisotopes and the Age of the Earth: Results of a Young-Earth Creationist Research Initiative, Volume II, L. Vardiman, A. Snelling, and E. Chaffin, eds., Institute for Creation Research, Santee, CA, 587-630, 2005.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Scaling laws for time-dependent stagnant lid convection in a spherical shell,” Phys. Earth Planet. Inter., 149, 361-370, 2005.
U. Walzer, R. Hendel, and J. Baumgardner, “Plateness of the oceanic lithosphere and the thermal evolution of the Earth's mantle,” in High Performance Computing in Science and Engineering '05, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3540283773, 289-304, 2005.
U. Walzer, R. Hendel, and J. Baumgardner, “The effects of a variation of the radial viscosity profile on mantle evolution,” Tectonophysics, 384, 55-90, 2004.
C. C. Reese, V. S. Solomatov, J. R. Baumgardner, D. R. Stegman, and A. V. Vezolainen, “Magmatic evolution of impact-induced Martian mantle plumes and the origin of Tharsis,” J. Geophys. Res., 109, 10.1029/2003 JE002222, 2004.
U. Walzer, R. Hendel, and J. Baumgardner, “Toward a geochemical model for the evolution of the Earth’s mantle,” in High Performance Computing in Science and Engineering '04, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-22943-4, 395-454, 2004.
U. Walzer, R. Hendel, and J. Baumgardner, “Generation of plate-tectonic behavior and a new viscosity profile of the Earth's mantle,” in NIC Symposium 2004 Proceedings, D. Wolf, G. Münster, and M. Kremer (eds.), NIC Series 20, 419-428., ISBN 3-00-012372-5, 2003.
J. R. Baumgardner, “Catastrophic plate tectonics: the physics behind the Genesis Flood,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., Editor, Creation Science Fellowship, Pittsburgh, PA, 113-126, 2003.
D. R. Humphreys, S. A. Austin, J. R. Baumgardner, and A. A Snelling, “Helium diffusion rates support accelerated nuclear decay,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 175-196, 2003.
L. Vardiman, S. A. Austin, J. R. Baumgardner, E. F. Chafin, D. B. DeYoung, D. R. Humphreys and A. A Snelling, “Radioisotopes and the age of the Earth,” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 337-348, 2003.
U. Walzer, R. Hendel, and J. Baumgardner, “Viscosity stratification and a 3-D compressible spherical shell model of mantle evolution”, in High Performance Computing in Science and Engineering '03, E. Krause, W. Jäger, and M. Resch, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-40850-9, 27-67, 2003.
D. R. Stegman, A.M. Jellinek, S. A. Zatman, J. R. Baumgardner, and M. A. Richards, “An early lunar core dynamo driven by thermochemical mantle convection,” Nature, 421, 143-146, 2003.
M. F. Horstemeyer and J. R. Baumgardner, “What initiated the Flood cataclysm?” in Proceedings of the Fifth International Conference on Creationism, R. L. Ivey, Jr., editor, Creation Science Fellowship, Pittsburgh, PA, 155-164, 2003.
J. R. Baumgardner, “Catastrophic plate tectonics: the geophysical context of the Genesis Flood,” “Dealing carefully with the data,” and “A constructive quest for truth,” all contributions to a “Forum on Catastrophic Plate Tectonics,” Ex Nihilo Technical Journal, 16, 57-85, 2002.
(http://www.answersingenesis.org/home/area/magazines/tj/docs/16n1p57Forum.asp)
H.-P. Bunge, M. A. Richards, and J. R. Baumgardner, “Mantle-circulation models with sequential data assimilation: inferring present-day mantle structure from plate-motion histories,” Phil. Trans. R. Soc. Lond. A., 360, 2545-2567, 2002.
D. A. Randall, T. D. Ringler, R. P. Heikes, P. Jones, and J. Baumgardner, “Climate Modeling with Spherical Geodesic Grids,” Computing in Science and Engineering, 4(5), 32-41, 2002.
U. Walzer, R. Hendel, and J. Baumgardner, “Variation of non-dimensional numbers and a thermal evolution model of the Earth's mantle,” in High Performance Computing in Science and Engineering '02, E. Krause and W. Jäger, eds., Springer-Verlag, Berlin Heidelberg New York, ISBN 3-540-43860-2, 89-103, 2002.
D. R. Stegman, M.A. Richards, and J.R. Baumgardner, Effects of depth dependent viscosity and plate motions on maintaining a relatively uniform MORB reservoir in whole mantle flow, J. Geophys. Res., 107, (B6), 2116, doi:10.1029/2001JB000192, 2002.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Survival of impact-induced thermal anomalies in the Martian mantle,” J. Geophys. Res.- Planets, 107(10), 5082-5092, 2002.
D. Majewski, D. Liermann, P. Prohl, B. Ritter, M. Buchhold, T. Hanisch, G. Paul, W. Wergen, and J. Baumgardner, “The global icosahedral-hexagonal grid point model GME: Operational version and high resolution tests,” Mon. Wea. Rev., 130, 319-338, 2002.
M. A. Richards, W.-S. Yang, and J. R. Baumgardner, “The role of a low viscosity zone in stabilizing plate tectonics: implications for comparative terrestrial planetology,” Geochem., Geophys., Geosys., 2, 2001.
J. R. Baumgardner, “Distribution of Radioactive Isotopes in the Earth” in Radioisotopes and the Age of the Earth: A Young Earth Creation Research Initiative, Larry Vardiman, Andrew A. Snelling, and Eugene F. Chaffin, eds., Institute for Creation Research, Santee, CA, 49-94, 2000.
J. K. Dukowicz and J. R. Baumgardner, “Incremental remapping as a transport/advection algorithm,” J. Comp. Phys., 160, 318-335, 2000.
W.-S. Yang and J. R. Baumgardner, "Matrix-dependent transfer multigrid method for strongly variable viscosity infinite Prandtl number thermal convection," Geophys. and Astrophys. Fluid Dyn., 92, 151-195, 2000.
H. R. Wenk, J. R. Baumgardner, C. N. Tome, and R. Lebensohn, "A deformation model to explain anisotropy of the inner core," J. Geophys. Res., 105, 5663-5677, 2000.
M. A. Richards, H.-P. Bunge, C. Lithgow-Bertelloni, and J. R. Baumgardner, "Mantle convection and plate motion history: Toward general circulation models," History and Dynamics of Global Plate Motions, AGU Monograph Series, 1999.
M. A. Richards, H.-P. Bunge, Y. Ricard, and J. R. Baumgardner, "Polar wandering in mantle convection models," Geophys. Res. Lett., 26, 1777-1780, 1999.
H.-P. Bunge, M. A. Richards, C. Lithgow-Bertelloni, J. R. Baumgardner, S. P. Grand, and B. A. Romanowicz, "Time scales and heterogeneity structure in geodynamic earth models," Science, 280, 91-95, 1998.
Hans-Peter Bunge, Mark A. Richards, and John R. Baumgardner, "A sensitivity study of 3-D spherical mantle convection at 108 Rayleigh number: effects of depth-dependent viscosity, heating mode, and an endothermic phase change," J. Geophys. Res.,102, B6, 11991-12007, 1997.
Hans-Peter Bunge, Mark A. Richards, and John R. Baumgardner, "The effect of depth-dependent viscosity on the planform of mantle convection," Nature, 379, 436-438, 1996.
Hans-Peter Bunge and John R. Baumgardner, "Mantle convection modeling on parallel virtual machines," Computers in Physics, 9, 207-215, 1995.
S. A. Austin, J. R. Baumgardner, D. R. Humphreys, A. A. Snelling, L. Vardiman, and K. P. Wise, "Catastrophic Plate Tectonics: A Global Flood Model of Earth History," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 609-621, 1994.
J. R. Baumgardner, "Computer Modeling of the Large-Scale Tectonics Associated with the Genesis Flood," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 49-62, 1994.
J. R. Baumgardner, "Runaway Subduction as the Driving Mechanism for the Genesis Flood," Proceedings of the Third International Conference on Creationism, Technical Symposium Sessions, R. E. Walsh, ed., Creation Science Fellowship, Inc., Pittsburgh, PA, 63-75, 1994.
D. B. Kothe, J. R. Baumgardner, J. H. Cerutti, B. J. Daly, K. S. Holian, E. M. Kober, S. J. Mosso, J. W. Painter, R. D. Smith, and M. D. Torrey, "PAGOSA: A massively-parallel, multi-material hydrodynamics model for three-dimensional high-speed flow and high-rate material deformation," Proceedings of the 1993 Conference on High Performance Computing: Grand Challenges in Computer Simulation, Society for Computer Simulation, 9-14, 1993.
John R. Baumgardner, "3-D numerical investigation of the mantle dynamics associated with the breakup of Pangea," in Flow and Creep in the Solar System: Observations, Modeling, and Theory, D. B. Stone and S. K. Runcorn, eds., NATO ASI Series C, Vol. 391, 207-224, 1993.
M. A. Moreno, G. Schubert, J. Baumgardner, M. G. Kivelson, and D. A. Paige, "Io's volcanic and sublimation atmospheres," Icarus, 93, 63-81, 1991.
John R. Baumgardner, "Application of supercomputers to 3-D mantle convection," in The Physics of the Planets, S. K. Runcorn, ed., John Wiley and Sons, 199-231, 1988.
J. Baumgardner, M. A. Moreno, G. Schubert, and M. G. Kivelson, "Two classes of volcanic eruptions and their corresponding atmospheres on Io," Bull. Am. Astr. Assoc., 19(3), 856, 1987.
John R. Baumgardner, "Three-dimensional treatment of convective flow in the earth's mantle," J. Stat. Phys., 39, 501-511, 1985.
John R. Baumgardner and Paul O. Frederickson, "Icosahedral discretization of the two-sphere," SIAM J. Numer. Anal., 22, 1107-1115, 1985.
Selected Conference Abstracts
D. R. Stegman, M. Jellinek, J. Baumgardner, and M. Richards, “A mantle dynamics perspective on generating an early lunar dynamo,” (abstract) Eos, Trans. Am. Geophys. Union, 84, (2003 Fall Meeting Supplement), 2003.
D. R. Humphreys, J. R. Baumgardner, A. A. Snelling, and S. A. Austin, “Recently measured helium diffusion rate for zircon suggests inconsistency with U-Pb age for Fenton Hill granodiorite,” (abstract) Eos, Trans. Am. Geophys. Union, 84, (2003 Fall Meeting Supplement), 2003.
J. R. Baumgardner, Y. Yamagishi, and D.R. Stegman, “Efficient visual data exploration in high-resolution global dynamical models of the Earth's mantle,” (abstract) Eos, Trans. Am. Geophys. Union, 83, (2002 Fall Meeting Supplement), 2002.
J. R. Baumgardner, “3D numerical modeling of the dynamics of Earth's mantle,” Am. Phys. Soc. Four Corners Annual Meeting, Las Cruces, New Mexico, 2001.
J. R. Baumgardner, W.-S. Yang, and M. A. Richards, “The role of a low viscosity zone in stabilizing a plate tectonics style of mantle convection,” (abstract) Eos, Trans. Am. Geophys. Union, 81, (2000 Fall Meeting Supplement), 2000.
J. R. Baumgardner and W.-S. Yang, “Earthlike mantle convection from relatively simple rheology,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F26, 1999.
M. A. Richards, W.-S. Yang, and J. R. Baumgardner, “The effectiveness of finite yield stress in obtaining platelike surface velocities,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F962, 1999.
D. R. Stegman, M. A. Richards, and J. R. Baumgardner, “A parallel implementation of Lagrangian tracers in TERRA,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F950, 1999.
C. C. Reese, V. S. Solomatov, and J. R. Baumgardner, “Impacts and the thermal evolution of Mars,” (abstract) Eos, Trans. Am. Geophys. Union, 80, (1999 Fall Meeting Supplement), F618, 1999.
John R. Baumgardner, Mark A. Richards, Woo-Sun Yang, and Carolina R. Lithgow-Bertelloni, "3-D Spherical Models of Plate Motion With Laterally Varying Rheology," (abstract) Eos, Trans. Am. Geophys. Union, 79, (1998 Fall Meeting Supplement), F911, 1998.
John R. Baumgardner and Woo-Sun Yang, "A finite element multigrid formulation for variable viscosity in 3-D spherical geometry," (abstract) Eos, Trans. Am. Geophys. Union, 77, (Fall Meeting Supplement), F750, 1996.
J. R. Baumgardner, "Thermal runaway in the mantle" (abstract) Eos, Trans. Am. Geophys. Union, 75, 687, 1994.
John Baumgardner, "3-D numerical investigation of the mantle dynamics associated with the breakup of Pangea," (abstract) Eos, Trans. Am. Geophys. Union, 73, 1992 Fall Meeting Abstract Volume, 576-577, 1992.
JULY 16, 2022
Creation Science Fellowship privately showed a movie to a small in-house audience about our stolen presidential election. This meeting was not broadcast or recorded.
July 2, 2022
Husband and wife team, Dan and Gay Zambrano gave the following presentation:
Addict Brain Normal Brain

About our speakers: Dr. Gay Zambrona is a 1991 graduate of the Ohio State University School of Veterinary Medicine. She fulfilled an externship at the London Zoo. She has been practicing full-time small animal and exotics medicine in the Long Beach/West Orange County area since 1991. She has also held part-time clinical and consulting positions in laboratory animal medicine for 15 years. Husband Dan Zambrano has a B.S. in Marine Biology from Cal State Long Beach. He is a Licensed Registered Veterinary Technician. He has worked at Cabrillo Aquarium, the Los Angeles Harbor Dept., California State Fish & Game Dept., and the L.A. Zoo before partnering in the mobile veterinary practice with his wife.

Both Dan and Gay are Ordained Pastors in the Free Methodist Church. They love teaching and preaching the Word of God to people of ALL AGES. They have worked with students from 4th & 5 th graders, through High School, College, and beyond!
June 18, 2022

With the kind permission of Former Military Space Engineer, Spike Psarris, we watched his newest expanded and updated edition of WHAT YOU AREN'T BEING TOLD ABOUT ASTRONOMY Volume I - OUR CREATED SOLAR SYSTEM. This visually stunning video shows example after example of recent discoveries among the planets and their moons that, according to conventional scientific wisdom, shouldn't be there. Scientific convention says the Solar System formed all by itself without a Creator, 4.5 billion years ago. But the heavenly bodies themselves display evidence that suggests otherwise.
To view trailers of Spike's Creation Astronomy videos, copy and paste the following into your url window: https://www.creationastronomy.com/videos/
To view trailers of Spike's Creation Astronomy videos, copy and paste the following into your url window: https://www.creationastronomy.com/videos/

About our Video Producer/Narrator:
Spike Psarris was previously an engineer in the United States military space program. He entered that program as an atheist and an evolutionist. He left it as a creationist and a Christian. In this video, he explains some of the evidence that caused that transformation.
Mr. Psarris is a Logos Research Associate.
To preview this and other Creation Astronomy videos, please visit www.creationastronomy.com.
Spike Psarris was previously an engineer in the United States military space program. He entered that program as an atheist and an evolutionist. He left it as a creationist and a Christian. In this video, he explains some of the evidence that caused that transformation.
Mr. Psarris is a Logos Research Associate.
To preview this and other Creation Astronomy videos, please visit www.creationastronomy.com.
June 4, 2022

We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 11 & 12 *
- Brilliant: Made in the Image of God
Ancient cultures reveal the rapid development of intelligent and creative people who were made in the image of God; not slow evolutionary advancement.
- A Matter of Time
The vast majority of dating methods reveal a recent creation and the impact of leaving this information out of student's education.
- Brilliant: Made in the Image of God
Ancient cultures reveal the rapid development of intelligent and creative people who were made in the image of God; not slow evolutionary advancement.
- A Matter of Time
The vast majority of dating methods reveal a recent creation and the impact of leaving this information out of student's education.

About our video speaker:
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
May 21, 2022
Joseph Kezele, M.D., gave a live, in-person presentation titled, "FROM DARWIN TO DICTATORSHIP." <<<(Double click)
This hard-hitting presentation is born from Dr. Kezele's personal experiences in Eastern Europe and opens with the “Laying the Foundation” for evolution that Ernst Häckel, laid down in Germany and in Europe as a whole. It then reviews how this foundation was used to promote racism and how the Nazis took advantage of that foundation. The scene then shifts to Soviet Russia after “Revolution of Foundation” and a recounting of some of the terrible events that occurred there in an evolutionary based political system. The last sections are “Evolution of Foundation” in America and “The Consequences”. They show why we now have to deal with racism, abortion, euthanasia, embryonic stem cells, and rationing of health care.
This hard-hitting presentation is born from Dr. Kezele's personal experiences in Eastern Europe and opens with the “Laying the Foundation” for evolution that Ernst Häckel, laid down in Germany and in Europe as a whole. It then reviews how this foundation was used to promote racism and how the Nazis took advantage of that foundation. The scene then shifts to Soviet Russia after “Revolution of Foundation” and a recounting of some of the terrible events that occurred there in an evolutionary based political system. The last sections are “Evolution of Foundation” in America and “The Consequences”. They show why we now have to deal with racism, abortion, euthanasia, embryonic stem cells, and rationing of health care.
Dr. Kezele's Slideshow:

About our speaker: Joseph Kezele, MD, studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomate of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians. He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, overseeing six creation science chapters throughout Arizona. He is Associate Professor at Arizona Christian University in Phoenix, teaching Human Genetics, Cell and Molecular Biology, Pathophysiology, Organic Chemistry and Creation Science Apologetics. Dr, Kezele has also been named a Logos Research Associate. He serves as Chairman of Deacons and teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona. He has participated in over 21 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, and Armenia. Dr. Kezele is one of the principle presenters in the Institute for Creation Research’s 4 DVD series, " Made in His Image". Four years ago, he returned to the Ukraine for three intense weeks of teaching Creation Science Apologetics and Evangelism as part of the activities prompted by the proclamation of the President of the Ukraine recognizing the 500th anniversary of Martin Luther's sparking the Reformation in Germany.
Dr. Kezele is a Logos Research Associate. He is also a Fellow of the Creation Research Society.
Dr. Kezele is a Logos Research Associate. He is also a Fellow of the Creation Research Society.
May 7, 2020
MOVIE NIGHT
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 9&10
- The Rocks Cry Out Curriculum Lesson 9: Explosive Evidence for Creation
Mt. St. Helens provides a model to understand some of the processes happening around the world during the Flood of Noah.
-The Rocks Cry Out Curriculum Lesson 10: Science starts with Creation
Consensus does not determine truth and not all scientists believe in evolution.
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 9&10
- The Rocks Cry Out Curriculum Lesson 9: Explosive Evidence for Creation
Mt. St. Helens provides a model to understand some of the processes happening around the world during the Flood of Noah.
-The Rocks Cry Out Curriculum Lesson 10: Science starts with Creation
Consensus does not determine truth and not all scientists believe in evolution.

About our video speaker:
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.

In CELEBRATION OF THE RESURRECTION OF JESUS CHRIST, the Creation Science Fellowship of Calvary Chapel Tustin forwent meeting on Easter Eve, Saturday night, April 16, 2022.
We pray you had a
HAPPY RESURRECTION DAY!
We pray you had a
HAPPY RESURRECTION DAY!
April 2, 2022

MOVIE NIGHT
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 7&8
- The Rocks Cry Out Curriculum Lesson 7: Science Is A Tool
The very laws of science support the idea of a Creator.
-The Rocks Cry Out Curriculum Lesson 8: The Grand Canyon
The Grand Canyon is a perfect example of the evidence supporting a worldwide flood.
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 7&8
- The Rocks Cry Out Curriculum Lesson 7: Science Is A Tool
The very laws of science support the idea of a Creator.
-The Rocks Cry Out Curriculum Lesson 8: The Grand Canyon
The Grand Canyon is a perfect example of the evidence supporting a worldwide flood.

About our video speaker:
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
MARCH 19, 2022

Monte Fleming, Ph.D. Geology, surveyed his new book, "STORIES ABOUT EARTH'S HISTORY: A Geologist's Dissent From Deep Time. "
Citing contents from his book, Dr. Fleming gave powerful geologic evidence that Earth's continents are YOUNG - only thousands of years old, not millions.
About our Speaker:
Monte Fleming holds a PhD in earth science. He is a teacher and a researcher, and his projects have included studies of whale fossils in Peru, analyses of giant rock basins on the tops of mesas in Arizona, and laboratory flume research (examining the deposit of sediments from rapidly flowing muddy water). He works at Loma Linda University in Loma Linda, California, where he lives with his wife, his oldest son, and his two-year-old triplets.
Citing contents from his book, Dr. Fleming gave powerful geologic evidence that Earth's continents are YOUNG - only thousands of years old, not millions.
About our Speaker:
Monte Fleming holds a PhD in earth science. He is a teacher and a researcher, and his projects have included studies of whale fossils in Peru, analyses of giant rock basins on the tops of mesas in Arizona, and laboratory flume research (examining the deposit of sediments from rapidly flowing muddy water). He works at Loma Linda University in Loma Linda, California, where he lives with his wife, his oldest son, and his two-year-old triplets.
MARCH 5, 2022

MOVIE NIGHT
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 5 & 6 *
- The Rocks Cry Out Curriculum Lesson 5: Dinosaurs and Dragons
Dinosaurs are used extensively to convince children that evolution is a fact and mankind evolved to appear upon the Earth millions of years AFTER dinosaurs went extinct. Yet cultures around the world and throughout history have references to these creatures. This lesson examines how dinosaurs fit into biblical history and reveals recent irrefutable discoveries confirming that dinosaurs were created quite recently – right alongside mankind.
-The Rocks Cry Out Curriculum Lesson 6: The Age of Creation
The belief in billions of years of Earth history implies that death, disease, bloodshed, and extinction have been around long before mankind appeared. If this were true, death is not really the penalty for mankind’s sin and God is the “author of death”. This lesson examines exactly how dating methods work, shows why they give erroneous results, and climaxes with a tour through a typical science and history museum which all too often are simply billion-dollar propaganda machines promoting non-biblical philosophical beliefs disguised as science.
We watched Bruce Malone's videos THE STONES CRY OUT, Lessons 5 & 6 *
- The Rocks Cry Out Curriculum Lesson 5: Dinosaurs and Dragons
Dinosaurs are used extensively to convince children that evolution is a fact and mankind evolved to appear upon the Earth millions of years AFTER dinosaurs went extinct. Yet cultures around the world and throughout history have references to these creatures. This lesson examines how dinosaurs fit into biblical history and reveals recent irrefutable discoveries confirming that dinosaurs were created quite recently – right alongside mankind.
-The Rocks Cry Out Curriculum Lesson 6: The Age of Creation
The belief in billions of years of Earth history implies that death, disease, bloodshed, and extinction have been around long before mankind appeared. If this were true, death is not really the penalty for mankind’s sin and God is the “author of death”. This lesson examines exactly how dating methods work, shows why they give erroneous results, and climaxes with a tour through a typical science and history museum which all too often are simply billion-dollar propaganda machines promoting non-biblical philosophical beliefs disguised as science.

About our video speaker:
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
Bruce spent 30 years working as a research leader for the Dow Chemical Corporation, has a degree in Chemical Engineering, and is responsible for key innovations which have resulted in 18 patents. But his passion is sharing the relevance and evidence for creation, so he retired early to become full time director of Search for the Truth Ministries. Bruce is an Ambassador with Logos Research Associates.
February 25, 2022

Jerry Bergman, Ph.D. (Human Biology); Ph.D. (Measurement and Evaluation); M.S.O.H. (Occupational Health); M.P.H. M.P.H. (Public Health); M.S.B.S. (Biomedical Science); M.A. (Social Psychology); M.Ed. (Counseling and psychology); B.S., (Sociology, Biology, Psychology); A.A. (Biology, Behavioral Science).
Dr. Bergman spoke on the topic of his newest book, co-authored with three other scientists, "APES AS ANCESTORS."
About our speaker:
(The following excerpt from another website -creation.com/dr-jerry-bergmanwebsite - is antiquated and in need of substantial updating. Dr. Bergman now has more than 100 publications to his name, including numerous books he has authored since the time this excerpt was posted.)
Dr. Bergman spoke on the topic of his newest book, co-authored with three other scientists, "APES AS ANCESTORS."
About our speaker:
(The following excerpt from another website -creation.com/dr-jerry-bergmanwebsite - is antiquated and in need of substantial updating. Dr. Bergman now has more than 100 publications to his name, including numerous books he has authored since the time this excerpt was posted.)

Jerry Bergman has taught biology, genetics, chemistry, biochemistry, anthropology, geology, and microbiology at Northwest State College in Archbold OH for over 25 years. He has 9 degrees, including 7 graduate (= ‘post-graduate’ in some non-US systems) degrees. Dr Bergman is a graduate of Medical College of Ohio, Wayne State University in Detroit, The University of Toledo, and Bowling Green State University. He has over 800 publications in 12 languages and 20 books and monographs. He has also taught at the Medical College of Ohio where he was a research associate in the department of experimental pathology, and he also taught 6 years at the University of Toledo, and 7 years at Bowling Green State University.
Among his books is a monograph on peer evaluation published by the College Student Journal Press, a Fastback on the creation-evolution controversy published by Phi Delta Kappa, a book on vestigial organs with Dr George Howe (‘Vestigial Organs’ are Fully Functional), a book on psychology and religious cults, a book on religious discrimination published by Onesimus Press, and a book on mental health published by Claudius Verlag in München. He has also published a college textbook on evaluation (Boston, Houghton Mifflin Co.), and has contributed to dozens of other textbooks. He was also a consultant for over 20 science text books, mostly biology and biochemistry.
Dr Bergman has presented over one hundred scientific papers at professional and community meetings in the United States, Canada, and Europe. To discuss his research, he has been a featured speaker on many college campuses throughout the United States and Europe, and is a frequent guest on radio and television programs. His research has made the front page in newspapers throughout the country, has been featured by the Paul Harvey Show several times, and has been discussed by David Brinkley, Chuck Colson, and other nationally known commentators on national television.
His other work experience includes over ten years experience at various Mental Health/Psychology clinics as a licensed professional clinical counselor and three years full time corrections research for a large county circuit court in Michigan and inside the walls of Jackson Prison (SPSM), the largest walled prison in the world. He has also served as a consultant for CBS News, ABC News, Reader’s Digest, Amnesty International, several government agencies and for two Nobel Prize winners, including the inventor of the transistor. In the past decade he has consulted or has testified as an expert witness or consultant in almost one-hundred court cases. A Fellow of the American Scientific Association, member of The National Association for the Advancement of Science, and many other professional associations, he is listed in Who’s Who in America, Who’s Who in the Midwest and in Who’s Who in Science and Religion.
* Subject to James 4:13-15 - " . . . For ye ought to say, If the Lord will, we shall live, and do this, or that."
Among his books is a monograph on peer evaluation published by the College Student Journal Press, a Fastback on the creation-evolution controversy published by Phi Delta Kappa, a book on vestigial organs with Dr George Howe (‘Vestigial Organs’ are Fully Functional), a book on psychology and religious cults, a book on religious discrimination published by Onesimus Press, and a book on mental health published by Claudius Verlag in München. He has also published a college textbook on evaluation (Boston, Houghton Mifflin Co.), and has contributed to dozens of other textbooks. He was also a consultant for over 20 science text books, mostly biology and biochemistry.
Dr Bergman has presented over one hundred scientific papers at professional and community meetings in the United States, Canada, and Europe. To discuss his research, he has been a featured speaker on many college campuses throughout the United States and Europe, and is a frequent guest on radio and television programs. His research has made the front page in newspapers throughout the country, has been featured by the Paul Harvey Show several times, and has been discussed by David Brinkley, Chuck Colson, and other nationally known commentators on national television.
His other work experience includes over ten years experience at various Mental Health/Psychology clinics as a licensed professional clinical counselor and three years full time corrections research for a large county circuit court in Michigan and inside the walls of Jackson Prison (SPSM), the largest walled prison in the world. He has also served as a consultant for CBS News, ABC News, Reader’s Digest, Amnesty International, several government agencies and for two Nobel Prize winners, including the inventor of the transistor. In the past decade he has consulted or has testified as an expert witness or consultant in almost one-hundred court cases. A Fellow of the American Scientific Association, member of The National Association for the Advancement of Science, and many other professional associations, he is listed in Who’s Who in America, Who’s Who in the Midwest and in Who’s Who in Science and Religion.
* Subject to James 4:13-15 - " . . . For ye ought to say, If the Lord will, we shall live, and do this, or that."
February 19, 2022

Jim Solliday, President of the Microscopical Society of Southern California, gave a presentation titled, "FOSSILS FROM AN OLD COAL MINE - Soft Tissue Structures from the Paleozoic"
About our speaker:
Jim Solliday was the 2019 recipient of the State Microscopical Society of Illinois (SMSI) 's August Köhler Award, the most prestigious honor given in the field of microscopy. SMSI published the following recognition of Jim's many accomplishments:
James D. Solliday is recognized for his numerous contributions to light microscopy and active participation in microscopical societies. He was a principal founder of the Microscopical Society of Southern California (MSSC) in 1996 and is currently the MSSC president, a chair he has held every year since 2002. He is also a long-time contributor to the Los Angeles Microscopical Society, and a member of the American Circuit of the Postal Microscopical Society, Quekett Microscopical Club, and the International Society for Diatom Research. Mr. Solliday is well known for his extensive knowledge of microscope history. He assists microscope collectors worldwide and was the curator of the Frank Lundy Antique Microscope Collection, which was publicly exhibited at the San Francisco international airport.
Mr. Solliday has taught workshops on diatoms, illumination methods, staining, photomicrography, microscope maintenance, and the history and identification of antique microscopes. He has published dozens of microscopy articles, including contributions to the MSSC, San Francisco Microscopical Society, Los Angeles Microscopical Society, and The Quekett Journal of Microscopy. He was instrumental in obtaining and donating surplus microscopes to area schools, and has been a regular visiting lecturer to local classrooms.
For 35 years, Mr. Solliday has operated his own business called Educational Photo Lab, which specializes in photomicrography. He has produced photomicrographs for Hollywood movies, documentaries, litigation exhibits, contamination analysis, authors, Ph.D. students, and publishers, with more than 450 microscope images published in textbooks. Beyond the micro world, he served for 31 years as a professional firefighter and medical technician for the city of Costa Mesa, CA.
About our speaker:
Jim Solliday was the 2019 recipient of the State Microscopical Society of Illinois (SMSI) 's August Köhler Award, the most prestigious honor given in the field of microscopy. SMSI published the following recognition of Jim's many accomplishments:
James D. Solliday is recognized for his numerous contributions to light microscopy and active participation in microscopical societies. He was a principal founder of the Microscopical Society of Southern California (MSSC) in 1996 and is currently the MSSC president, a chair he has held every year since 2002. He is also a long-time contributor to the Los Angeles Microscopical Society, and a member of the American Circuit of the Postal Microscopical Society, Quekett Microscopical Club, and the International Society for Diatom Research. Mr. Solliday is well known for his extensive knowledge of microscope history. He assists microscope collectors worldwide and was the curator of the Frank Lundy Antique Microscope Collection, which was publicly exhibited at the San Francisco international airport.
Mr. Solliday has taught workshops on diatoms, illumination methods, staining, photomicrography, microscope maintenance, and the history and identification of antique microscopes. He has published dozens of microscopy articles, including contributions to the MSSC, San Francisco Microscopical Society, Los Angeles Microscopical Society, and The Quekett Journal of Microscopy. He was instrumental in obtaining and donating surplus microscopes to area schools, and has been a regular visiting lecturer to local classrooms.
For 35 years, Mr. Solliday has operated his own business called Educational Photo Lab, which specializes in photomicrography. He has produced photomicrographs for Hollywood movies, documentaries, litigation exhibits, contamination analysis, authors, Ph.D. students, and publishers, with more than 450 microscope images published in textbooks. Beyond the micro world, he served for 31 years as a professional firefighter and medical technician for the city of Costa Mesa, CA.
February 5, 2022
Drs. Steve and Kelly Austin gave a ZOOM presentation titled, "THE INCREDIBLE MIND OFTHE HONEY BEE."
About our speakers:
January 15, 2022

While recovering from a light bout with COVID-19, Jim Pamplin gave a presentation by ZOOM only titled,
"PRECEPTS IN NATURAL THEOLOGY" . No in-house meeting of the Creation Science Fellowship took place.
About our speaker:
Jim Pamplin led the Creation Science Fellowship of Calvary Chapel Costa Mesa for 18 years before moving the Fellowship to Calvary Chapel Tustin, more than a year ago. Jim is a co-founder and director of Logos Research Associates, a fellowship of more than 70 scientists and scholars dedicated to the proposition that "Good Science Affirms Scripture." Jim is a missionary of Calvary Chapel Costa Mesa, ministering to world culture in the area of Creation Apologetics.
"PRECEPTS IN NATURAL THEOLOGY" . No in-house meeting of the Creation Science Fellowship took place.
About our speaker:
Jim Pamplin led the Creation Science Fellowship of Calvary Chapel Costa Mesa for 18 years before moving the Fellowship to Calvary Chapel Tustin, more than a year ago. Jim is a co-founder and director of Logos Research Associates, a fellowship of more than 70 scientists and scholars dedicated to the proposition that "Good Science Affirms Scripture." Jim is a missionary of Calvary Chapel Costa Mesa, ministering to world culture in the area of Creation Apologetics.
January 1, 2022
The Creation Science Fellowship did not meet on the first Saturday of January, 2022, because it fell on New Years Day.
December 2021
Calvary Chapel Tustin honored the Lord's birth with a flurry of special events and activities throughout the month of December. Supplemental ministries took a vacation during December to facilitate special programming for the month.
The Creation Science Fellowship did not meet during the month of December, 2021.
The Creation Science Fellowship did not meet during the month of December, 2021.
NOVEMBER 13, 2021
(Due to a facility maintenance event, the CSF meeting originally scheduled for November 6 was postponed until November 13)

Ross Anderson, Ph.D., Biochemistry, gave a LIVE, IN-PERSON presentation titled, "HEMOGLOBIN: EVIDENCE OF DESIGN."
About our speaker:
Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society.
About our speaker:
Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society.
OCTOBER 16, 2021

David Dana-Bashan gave* a LIVE, IN-PERSON presentation titled '16 NEARLY IMPOSSIBLE ISSUES FOR EVOLUTIONISTS TO ANSWER.' David argues, " Any solutions that evolutionists propose to such issues, inevitably and invariably violate basic, well-known laws of mathematics, physics, and chemistry."
About our speaker: Mr. Dana-Bashan earned a B.S. in Mathematics, an M.S. in Mathematics, and a B.S. in Chemistry (A.C.S. Certified), all from Purdue University. He is a member of the honorary chemistry fraternity Phi Lambda Upsilon, and has a lifetime of experience in the hard sciences. David is also one of the five dozen persons in the world who have been identified as possessing Highly Superior Autobiographical Memory. Give him the date of any day in history, and he can tell you on what day of the week it occurred and relate that day to events of the time. For dates in his own lifetime, he can tell you news headline items of that day, week, and month, and what songs were popular on the radio.
About our speaker: Mr. Dana-Bashan earned a B.S. in Mathematics, an M.S. in Mathematics, and a B.S. in Chemistry (A.C.S. Certified), all from Purdue University. He is a member of the honorary chemistry fraternity Phi Lambda Upsilon, and has a lifetime of experience in the hard sciences. David is also one of the five dozen persons in the world who have been identified as possessing Highly Superior Autobiographical Memory. Give him the date of any day in history, and he can tell you on what day of the week it occurred and relate that day to events of the time. For dates in his own lifetime, he can tell you news headline items of that day, week, and month, and what songs were popular on the radio.
october 2, 2021

Joseph Kezele, MD, gave a two-part LIVE Presentation lecture, titled: VALUBLE AND VISCIOUS VIRUSES AND A VACCINE, (Parts 1 & 2). Part 1 introduced the subject of viruses in general, and Part 2 focused on specifics of COVID-19.
To view these recordings, please go to https://drive.google.com/drive/folders/11foCdXmOp__vznxJn2iwfTRJ1ls8rSL9.
Select 10022021A for Part 1
Select 10022021B for Part 2
To view these recordings, please go to https://drive.google.com/drive/folders/11foCdXmOp__vznxJn2iwfTRJ1ls8rSL9.
Select 10022021A for Part 1
Select 10022021B for Part 2

About our speaker: Joseph Kezele, MD, studied the sciences and music at the University of Arizona, while earning his B.A. in Russian in 1971, and his M.D. in 1975. He then completed a Pediatric Residency and practiced Emergency Medicine for 25 years in the Phoenix area, and was a diplomate of the American Board of Emergency Medicine and fellow of the American College of Emergency Physicians. He is now retired from the practice of medicine, serves as President of the Arizona Origin Science Association, overseeing six creation science chapters throughout Arizona. He is Associate Professor at Arizona Christian University in Phoenix, teaching Human Genetics, Cell and Molecular Biology, Pathophysiology, Organic Chemistry and Creation Science Apologetics. Dr, Kezele has also been named a Logos Research Associate. He serves as Chairman of Deacons and teaches Creation Science Evangelism at Black Mountain Baptist Church in Cave Creek, Arizona. He has participated in over 21 foreign mission trips, teaching medicine, standard evangelism and creation science evangelism in India, Zambia, Mexico, Russia, Latvia, Lithuania, the Ukraine, Bosnia-Herzegovina, and Armenia. Dr. Kezele is one of the principle presenters in the Institute for Creation Research’s 4 DVD series, " Made in His Image". Four years ago, he returned to the Ukraine for three intense weeks of teaching Creation Science Apologetics and Evangelism as part of the activities prompted by the proclamation of the President of the Ukraine recognizing the 500th anniversary of Martin Luther's sparking the Reformation in Germany.
Dr. Kezele is a Logos Research Associate.
Dr. Kezele is a Logos Research Associate.
September 18, 2021

We watched dvd Disc 1 (127 minutes) of the "RATE PREMIER CONFERENCE, Thousands . . . Not Billions" held on November 5, 2005.
The November 5, 2005 conference, held at Shadow Mountain Church in San Diego, (and the attending two-volume book Radioisotopes and the Age of the Earth), culminated 8 years of research by eight leading scientists, revealing multiple lines of evidence that earth experienced one or more episodes of accelerated radioactive decay in its past.
We will watch* dvd Disc 2 at a later date.
* If the Lord will . . . (James 4:13-15)
september 4, 2021
We watched threeYouTube videos related to the laying down of the geologic record, with an eye toward understanding the role of Noah's Flood.
These were :
EXPERIMENTS IN STRATIFICATION (https://www.youtube.com/watch?v=wFST2C32hMQ&t=47s)
These were :
EXPERIMENTS IN STRATIFICATION (https://www.youtube.com/watch?v=wFST2C32hMQ&t=47s)
Collectively, these videos challenge the simplistic interpretation of geological strata as have being laid down slowly in calm water over millions of years. The tired old mantra, "The present is the key to the past" FAILS. Rock layers give evidence of fast and recent catastrophic deposition on a global scale. These videos suggest that most strata were laid down rapidly by vast sheets of fast flowing water, loaded with sediments, which on such a massive scale, have no counterpart in the world today. Rock layers around the world give silent testimony to a recent, global, catastrophic flood.
AUGUST 21, 2021

Ross Anderson, Ph.D., Biochemistry, gave a LIVE, IN-PERSON presentation titled, FORETHOUGHT AND PLANNING IN THE DEVELOPMENT OF THE VERTEBRATE EYE: A Case Study for Intelligent Design (https://drive.google.com/drive/folders/11foCdXmOp__vznxJn2iwfTRJ1ls8rSL9)
About our speaker:
Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society.
About our speaker:
Dr. Anderson’s expertise is in the area of biochemistry and molecular biology. He has taught Biochemistry and helped to direct research projects of graduate and medical students at Baylor College of Medicine, Houston, TX. Dr. Anderson was a post-doctoral researcher in the Molecular Genetics Division of the Department of Ophthalmology at the Houston Neurosensory Center.
Dr. Anderson was a member of both the undergraduate and graduate faculty at Lamar University, Beaumont, TX. There he taught and directed the research activities of undergraduates and Masters of Science degree candidates in Biology.
Dr. Anderson’s research interests include structure-function studies of DNA polymerizing enzymes and the synthesis and expression of synthetic human genes in bacterial hosts. He has authored or co-authored several publications in major, peer-reviewed journals. He is a member of the American Chemical Society and Sigma Xi Research Society.
August 7, 2021

Matthew Cserhati, Ph.D. Bioinformatics, gave a LIVE, IN-PERSON presentation titled, CREATION SCIENCE ANALYSIS OF RECENT HUMAN FOSSILS. (https://drive.google.com/drive/folders/11foCdXmOp__vznxJn2iwfTRJ1ls8rSL9)
About our speaker (The following information is borrowed from Creation Ministries International's website at https://creation.com/dr-matthew-cserhati-cv.)
Education
During his Ph.D. studies, Matthew developed a transcription factor binding site dyad prediction algorithm, with application to crop species at the Biological Research Center in Szeged, Hungary. He also developed a dedicated website for housing haplotype information on candidate drought genes in barley and taught two science courses.
From 2010–2011, he took part in the development of the program OPTIMA in the laboratory of Chemical Kinetics in the Institute of Chemistry at ELTE University, in Budapest. This work involved the estimation of a number of Arrhenius parameters in a number of combustion reactions. He also taught Linux programming as a special course.
From 2011–2013, he worked as a post-doctoral research fellow at the University of Nebraska, Lincoln. The two main projects he was involved with included the analysis of Next-gen sequencing data to discover differentially expressed genes in the green algae, Chlamydomonas reinhardtii due to low carbon concentrations, and the analysis of the regulation of 177 ribosomal protein genes across ~30,000 experiments, representing hundreds of gigabytes of data from the ENCODE project.
From 2013–2017, he worked as a bioinformatics programmer at the University of Nebraska Medical Center. His tasks included the development of the NNTC-DCC database, a browsable, integrated website for storing experimental and clinical data on neuroAIDS patients. He was also involved in the demultiplexing, storage, and analysis of Next-gen sequencing data for wet-lab researchers. Other notable projects included the de novo genome assembly and annotation of ten NCLDV (NucleoCytoplasmic Large DNA Viruses), as well as the application of a motif prediction algorithm to the genomes of modern and archaic humans. He also helped teach a couple courses in bioinformatics.
In 2017 he started an M.A. program at Greenville Presbyterian Theological Seminary.
From 2018–2019, he took part in the prediction and analysis of differentially expressed genes in hepatocellular carcinoma (HCC) in mouse and human at the University of Texas Health Science Center at San Antonio.
From 2010 He performed over 200 translations and interpretation jobs in Hungarian, German, Swedish, Dutch and Norwegian. He was also a member of the American Translators Association and the Nebraska Association for Translators and Interpreters for several years and he led an informal biblical languages club at Grace Orthodox Presbyterian Church in San Antonio, Texas.
In 2001 he helped establish a creation science group in Hungary called the Protestant Creation Research Group. In 2015 he became a member of the Creation Research Society.
Creationist publications and articles (using several pseudonyms)
About our speaker (The following information is borrowed from Creation Ministries International's website at https://creation.com/dr-matthew-cserhati-cv.)
Education
- 2011, University of Szeged, Hungary, Ph.D., Bioinformatics
- 2010, University of Szeged, Hungary, B.Sc., Software Development
- 2003, Eötvös Loránd University, M.Sc., Biology
- Native Hungarian speaker
- Swedish, Advanced level, complex, oral
- Dutch, Intermediate level, oral + written
- German, Intermediate level, oral
- Norwegian, Intermediate level, written
- Beginning Greek, Greenville Presbyterian Theological Seminary
- Hebrew I+II, Greenville Presbyterian Theological Seminary
During his Ph.D. studies, Matthew developed a transcription factor binding site dyad prediction algorithm, with application to crop species at the Biological Research Center in Szeged, Hungary. He also developed a dedicated website for housing haplotype information on candidate drought genes in barley and taught two science courses.
From 2010–2011, he took part in the development of the program OPTIMA in the laboratory of Chemical Kinetics in the Institute of Chemistry at ELTE University, in Budapest. This work involved the estimation of a number of Arrhenius parameters in a number of combustion reactions. He also taught Linux programming as a special course.
From 2011–2013, he worked as a post-doctoral research fellow at the University of Nebraska, Lincoln. The two main projects he was involved with included the analysis of Next-gen sequencing data to discover differentially expressed genes in the green algae, Chlamydomonas reinhardtii due to low carbon concentrations, and the analysis of the regulation of 177 ribosomal protein genes across ~30,000 experiments, representing hundreds of gigabytes of data from the ENCODE project.
From 2013–2017, he worked as a bioinformatics programmer at the University of Nebraska Medical Center. His tasks included the development of the NNTC-DCC database, a browsable, integrated website for storing experimental and clinical data on neuroAIDS patients. He was also involved in the demultiplexing, storage, and analysis of Next-gen sequencing data for wet-lab researchers. Other notable projects included the de novo genome assembly and annotation of ten NCLDV (NucleoCytoplasmic Large DNA Viruses), as well as the application of a motif prediction algorithm to the genomes of modern and archaic humans. He also helped teach a couple courses in bioinformatics.
In 2017 he started an M.A. program at Greenville Presbyterian Theological Seminary.
From 2018–2019, he took part in the prediction and analysis of differentially expressed genes in hepatocellular carcinoma (HCC) in mouse and human at the University of Texas Health Science Center at San Antonio.
From 2010 He performed over 200 translations and interpretation jobs in Hungarian, German, Swedish, Dutch and Norwegian. He was also a member of the American Translators Association and the Nebraska Association for Translators and Interpreters for several years and he led an informal biblical languages club at Grace Orthodox Presbyterian Church in San Antonio, Texas.
In 2001 he helped establish a creation science group in Hungary called the Protestant Creation Research Group. In 2015 he became a member of the Creation Research Society.
Creationist publications and articles (using several pseudonyms)
- Cserhati, M., Baraminology data filtering method based on entropy measurement and its application in dinosaur and cephalopod data sets, Journal of Creation 33(3):55-65, 2019.
- Cserhati, M., Alquist, J., Baraminology suggests cryptic relationships among Caprimulgiformes, Journal of Creation 33(3):66-76, 2019.
- Cserhati, M., and Tay, J., Comparison of morphology-based and genomics-based baraminology methods. Journal of Creation 33(3):49-55, 2019.
- Cserhati, M., The green algae Chlamydomonas reinhardtii finds safety in numbers by design, J Creation 33(2):93-98, 2019.
- Cserhati, M., and Chira, A., Critique of the latest evolutionary models of the beginning of life. Journal of Creation 33(1):78–84, 2019.
- O’Micks, J., Baraminic analysis of archaic and modern human genomes. Journal of Creation 32(3):19–24, 2018.
- Cserhati, M., Swinging too far to the other side. Journal of Creation 32(2):60–62, 2018.
- Cserhati, M., What does the catechism of the Roman Catholic Church say about creation? Journal of Creation 32(2):79–82, 2018.
- Cserhati, M., and Shipton, W., The irreducibly complex ribosome is a unique creation in the three domains of life. Journal of Creation 31(2):94-102, 2017.
- O’Micks, J., Baraminic analysis of nucleocytoplasmic large DNA viruses. Journal of Creation 31(1):73-79, 2017.
- O’Micks, J., Creation perspective of nucleocytoplasmic large DNA viruses. Journal of Creation 30(3):110-117, 2016.
- O’Micks, J., Promoter evolution is impossible by random mutations. Journal of Creation 30(2):60-66, 2016.
- O’Micks, J., Pseudogenes and bacterial genome decay. Journal of Creation 30(1):104-110, 2016.
- O’Micks, J., Cnidarians turn evolutionary theory into jelly. Journal of Creation 29(3):71-79, 2015. See also the response in Journal of Creation 30(2):51, 2016.
- O’Micks, J., Bacterial genome decay from a baraminological viewpoint. Journal of Creation 29(2):122-130, 2015.
- Savanne, D., Denisovans menace evolution—a new chapter in the human origins debate. Journal of Creation 28(3):5-8, 2014.
- O’Micks, J., Small genome size of Utricularia gibba problematic for evolution but not creation. Journal of Creation 27(3):4-6, 2013.
- Arneigh, M.R., It’s a small world—microRNA cuts evolution down to size. Journal of Creation 27(2):85-90, 2013.
- Shan, L.E., Transposon amplification in rapid intrabaraminic diversification. Journal of Creation 23(2):110-117, 2009.
- Cserháti, M., Creation aspects of conserved non-coding sequences. Journal of Creation 21(2):101-108, 2007.
- O’Micks, J., A preliminary cephalopod baraminology study based on the analysis of mitochondrial genomes and morphological characteristics. Answers Research Journal 11:193–204, 2018.
- O’Micks, J., Further evidence that Homo naledi is not a member of the human holobaramin based on measurements of vertebrae and ribs. Answers Research Journal 10:103–113, 2017.
- O’Micks, J., Likely Discontinuity Between Humans and Non-Human Hominins Based on Endocranial Volume and Body Mass With a Special Focus on Homo naledi—A Short Analysis. Answers Research Journal 10:241-243, 2017.
- O’Micks, J., Rebuttal to “Reply to O’Micks concerning the geology and taphonomy of the Homo naledi site” and “Identifying humans in the fossil record: a further response to O’Micks”. Answers Research Journal 10:63–70, 2017.
- O’Micks, J., Reply to “Taxon sample in hominin baraminology: a response to O’Micks”. Answers Research Journal 9:373–375, 2016.
- O’Micks, J., Molecular structures shared by prokaryotes and eukaryotes show signs of only analogy and not homology. Answers Research Journal 9:284–292, 2016.
- O’Micks, J., Homo naledi probably not part of the human holobaramin based on baraminic re-analysis including postcranial evidence. Answers Research Journal 9:263–272, 2016.
- O’Micks, J., Baraminology classification based on gene content similarity measurement. Creation Research Society Quarterly 54:27–37, 2017.
- Yaugh, A., Baraminological analysis of a set of Archaea species based on genomic data. Creation Research Society Quarterly 53:140–154, 2016.
- O’Micks, J., Preliminary baraminological analysis of Homo naledi and its place within the human baramin. Journal of Creation Theology and Science Series B: Life Sciences 6: 31–39, 2016.
- More of Dawkins’ same old tired rhetoric. Review of Outgrowing God by Richard Dawkins
- O’Micks, J., Taking the bite out of evolution — critical review of Evolution’s Bite by Peter S. Ungar. Journal of Creation 31(3):24-27, 2017.
- O’Micks, J., Modern science catches up with Neandertal man, a review of The Neanderthals Rediscovered—How modern science is rewriting their story by Dimitra Papagianni and Michael A. Morse. Journal of Creation 32(1):38–42, 2017.
- O’Micks, J., and Clarey, T., Book Review of Almost Human, the astonishing tale of Homo naledi and the discovery that changed our human story, by Lee Berger and John Hawks. Answers Research Journal 10:187–194, 2017.
- Cserhati, M., Xiao, P., Guda, C., K-mer-Based Motif Analysis in Insect Species across Anopheles, Drosophila, and Glossina Genera and Its Application to Species Classification, Computational and Mathematical Methods in Medicine vol. 2019, 2019.
- Cserhati, M., Mooter, M.E., Fan, M., Wicks, B., Xiao, P., Pauley, M., and Guda, C., Motif comparison between human, Neandertals and Denisova. BMC Genomics 19(1):472, 2018 | doi:10.1186/s12864-018-4710-1.
- Lopez, W., Page, A.M., Carlson, D.J., Ericson, B.L., Cserhati, M.F., Guda, C., and Carlson, K.A., Analysis of immune-related genes during Nora virus infection of Drosophila melanogaster using next generation sequencing. AIMS Microbiology 4(1):123–139, 2018. doi: 10.3934/microbiol.2018.1.123.
- Cserhati, M., Motif content comparison between monocot and dicot species. Genom Data 3:128–36, 2015 | doi:10.1016/j.gdata.2014.12.006.
- Pettkó-Szandtner, A., Cserháti, M., Barrôco, R.M., Hariharan, S., Dudits, D., and Beemster, G.T., Core cell cycle regulatory genes in rice and their expression profiles across the growth zone of the leaf. J Plant Res 128(6):953—74, 2015 | doi:10.1007/s10265-015-0754-3.
- Cserhati, M., Pandey, S., Beaudoin, J.J., Baccaglini, L., Guda, C., and Fox, H.S., The National NeuroAIDS Tissue Consortium (NNTC) Database: an integrated database for HIV-related studies. Database (Oxford) 2015:bav074 | doi:10.1093/database/bav074.
- Szécsényi, M., Cserháti, M., Zvara, Á., Dudits, D., and Györgyey, J., Monitoring of transcriptional responses in roots of six wheat cultivars during mild drought stress. Cereal Research Communications 41(4):528–538, 2013 | doi:10.1556/CRC.41.2013.4.3.
- Cserháti, M., Turóczy, Z., Dudits, D., and Györgyey, J., The rice word landscape: a detailed catalogue of the rice motif content in the non-coding regions. OMICS 16(6):334–42, 2012. doi:10.1089/omi.2011.0056.
- Cserhati, M., Prommute – A promoter mutation simulation for modeling the evolution of genetic regulatory elements. Journal of Computer Science and Systems Biology 5:74–80, 2012 |doi:10.4172/jcsb.1000093.
- Brueggeman, A.J., Gangadharaiah, D.S., Cserhati, M., Diaz-Cano, D.C., Weeks, D.P., and Ladunga, I., Activation of the carbon concentrating mechanism by CO2 deprivation coincides with massive transcriptional restructuring in Chlamydomonas reinhardtii. Plant Cell 24(5):1860–75, 2012 | doi:10.1105/tpc.111.093435.
- Zsély, I.,Gy., Varga, T., Nagy, T., Cserháti, M., Turányi, T., Peukert, S., Braun-Unkhoff, M., Naumann, C., and Riedel, U., Determination of rate parameters of cyclohexane and 1-hexene decomposition reactions. Energy 43(1):85–93, 2012 | doi:10.1016/j.energy.2012.01.004.
- Turányi, T., Nagy, T., Zsély, I.Gy., Cserháti, M., Varga, T., Szabó, B.T., Sedyó, I., Kiss, P.T., Zempléni, A., and Curran, H.J, Determination of rate parameters based on both direct and indirect measurements. Int J Chem Kinet 44(5):284–302, 2012 | doi:10.1002/kin.20717.
- Áy, Z., Mihály, R., Cserháti, M., Kótai, É., and Pauk, J., The effect of high concentrations of glufosinate ammonium on the yield components of transgenic spring wheat (Triticum aestivum L.) constitutively expressing the bar gene. The Scientific World Journal 2012:657945 | doi:10.1100/2012/657945.
- Cseri, A., Cserháti, M., von Korff, M., Bettina, N., Gábor, V.H., Palágyi, A., Pauk, J., Dudits, D., and Törjék, O., Allele mining and haplotype discovery in barley candidate genes for drought tolerance. Euphytica 181:341, 2012 | doi:10.1007/s10681-011-0445-7.
- Cserháti, M., Turóczy, Z., Zombori, Z., Cserzo, M., Dudits, D., Pongor, S., and Györgyey, J., Prediction of new abiotic stress genes in Arabidopsis thaliana and Oryza sativa according to enumeration-based statistical analysis. Mol Genet Genomics 285(5):375–91, 2011 | doi:10.1007/s00438-011-0605-4.
- Turóczy, Z., Kis, P., Török, K., Cserháti, M., Lendvai, A., Dudits, D., and Horváth, G.V., Overproduction of a rice aldo-keto reductase increases oxidative and heat stress tolerance by malondialdehyde and methylglyoxal detoxification. Plant Mol Biol 75(4–5):399–412, 2011 | doi:10.1007/s11103-011-9735-7.
- Cserhati, M., and Gyorgyey, J., “Gene research in silico”, in The science of wheat breeding (ed. Denes Dudits), Winter Fair Ltd., Eger, 2006.
- Dudits, D., Cserháti, M., Miskolczi, P., and Horváth, G., “The growing family of plant cyclin dependent kinases with multiple functions in cellular and developmental regulation”, in Cell cycle control and development (ed. Dirk Inzé), Blackwell Publishing, Oxford, 2006.
- Baraminology suggests cryptic relationships among Caprimulgiformes with Jon Ahlquist. From Journal of Creation 33(3), 2019.
- Baraminology data filtering method based on entropy measurement and its application in dinosaur and cephalopod data sets from Journal of Creation 33(3), 2019.
- Comparison of morphology-based and genomics-based baraminology methods with Joel Tay. From Journal of Creation 33(3), 2019.
- One for all and all for one from Creation magazine 41(1) 2020.
- The wonderful world of bats from Creation magazine 41(1) 2020.
- The wonderfully designed cell cycle from Creation magazine 41(4) 2019.
- Evidence that modern humans and ‘super-archaic’ humans may have shared DNA
- What should Christians think about artificial selection and genetic modification?
- Jacob’s livestock
- More of the moth myth!
- Owls—masters of the night sky from Creation magazine 42(3) 2020.
- The priests of Darwin want your children
- Cosmic Biology with Paul Price
- Is the RubisCO enzyme an ineffective leftover of evolution?
- Snakes losing legs is not evolution!
- Do koalas prove that humans got part of their DNA from viruses?
- Racial mixing is perfectly biblical!
- Is a brain a human being?
- Weird synthetic proteins are no help to evolution. Changing the genetic code points to design
- What is Homo luzonensis?
- Evolution of middle ear bones in mammals from jaw bones in reptiles?
- Yale university professor renounces Darwinism!
- Reformation wall vandalized yet again
- From a rib to a 206-bone human
- Can mutations lead to new genetic information?
- Is the Shroud of Turin authentic?
- Researching creation after secular academia
- Rails derail evolution
- Ambopteryx – bird or bat-winged reptile?
- New sugar transport gene evolved in yeast?
- Were stem cell-like organisms the first forms of life?
- Did the ear bones of mammals really evolve from the jawbones of reptiles?
- Homeschool conference: great encouragement and some concerns
- Hachimoji DNA argues against evolution, despite recent claims
- Newly discovered Medusavirus turns evolutionary theory to stone
- Have astrobiologists really found a super-Earth?
- What is Archaeotherium? Pig or hippopotamus? What is Onychonycteris? Shrew or bat?
- More of Dawkins’ same old tired rhetoric - Review of Outgrowing God by Richard Dawkins
July 17, 2021

We watched a video titled, THE ALTERNATIVE: Creation's Competitive Edge by Dr. Rob Carter. (https://www.youtube.com/watch?v=ute0VkpNcd8)
About the video's featured speaker:
Dr Robert W. Carter, Speaker/senior scientist, Creation Ministries International (USA)
Education
Human genetic history
About the video's featured speaker:
Dr Robert W. Carter, Speaker/senior scientist, Creation Ministries International (USA)
Education
- 2003 University of Miami, Ph.D. Marine Biology
- 1992 Georgia Institute of Technology, B.S. Applied Biology
- Between my undergraduate and graduate work, I spent four years teaching high school science at a large college-prep school in NW Georgia. In addition to my teaching load (including AP biology, chemistry, physics, and electronics), I spent the winter months coaching the swimming and diving teams and the summer months running the outdoor high adventure program.
- In 1996, I was awarded the three-year Maytag doctoral fellowship by the University of Miami. When that expired, I received a one-year fellowship from the Institute for Marine Science. While working on my PhD, I designed and performed many experiments in marine ecology and genetic engineering and helped to develop new protocols for the rapid cloning of fluorescent protein genes. The green and red fluorescent proteins my coworker and I cloned from hard and soft corals were used to create transgenic zebrafish. We patented one of these protein genes and licensed it to Promega, Inc. under the trade name ‘Monster Green’.
- From 2001–2004, I helped design and build an aquaculture facility for Caribbean corals at UM’s Experimental Fish Hatchery. During these years I also performed over 500 research dives on the shallow coral reefs of the Florida Keys and Bahamas. Many of these were done at night to study the mass coral spawning episodes that happen at specific times during the warm summers.
- I spent two years after obtaining my PhD working for an engineering company, mainly focused on impact mitigation for the Key West Harbor dredging project (since the channel runs right through the coral reef).
- Upon leaving Miami, I was hired by the Institute for Creation Research to help on their GENE project. While there, I wrote computer programs to analyze human genetic data and managed to get one publication on this work into the secular literature.
- In 2006, I was hired by Creation Ministries International as a scientist, speaker, and writer.
- Carter RW, Sanford, JC (2012) A new look at an old virus: mutation accumulation in the human H1N1 influenza virus since 1918. Theoretical Biology and Medical Modelling 9:42 | doi:10.1186/1742-4682-9-42.
- Carter RW (2007) Mitochondrial diversity within modern human populations. Nucleic Acids Research 35(9):3039–3045.
- Gibbs PDL, Carter RW, and Schmale MC (2007) Fluorescent Proteins from Aquatic Species. US Patent #7,291,711.
- Carter RW, Schmale MS, and Gibbs PDL (2004) Cloning of anthozoan fluorescent protein genes. Comparative Biochemistry and Physiology, Part C 138:259–270.
- Carter RW (2003) Cnidarian Fluorescent Proteins. PhD Dissertation. University of Miami.
- Manica A, Carter RW (2000) Morphological and fluorescence analysis of the Montastraea annularis species complex in Florida. Marine Biology 137:899–906.
- Bad arguments for the Masoretic text
- Were the Egyptian pyramids built before the Flood?
- The inspiration of Scripture comes in various forms
- Is the Shroud of Turin authentic?
- Iron sharpening iron: the MT-LXX debate as a case study of Christian disagreement
- The Masoretic text of Genesis 5 and 11 is still the most reliable
- Is the Septuagint a superior text for the Genesis genealogies?
- Mary: the biblical woman behind the cultural legend
- Where was Eden? Part 1—examining pre-Flood geographical details in the biblical record
- Where was Eden? Part 2—geological considerations—examining pre-Flood geographical details in the biblical record
- Where was Eden?
- The genetic history of the Israelite nation
- Extensive mixing among Israelites and non-Israelites in biblical history
- Reply to James Denning on Extensive mixing among Israelites and non-Israelites
- Textual traditions and biblical chronology
- The biblical minimum and maximum age of the earth
- Reply to David Austin on The biblical minimum and maximum age of the earth
- How old was Cain when he killed Abel?
- Robert Carter gets everything wrong? Responding to even more ridiculous aspersions
- Genetic Entropy and the Gospel: Biological clocks say the earth is young!
- Responding to supposed refutations of genetic entropy from the ‘experts’ (with Paul Price and comments by Dr John Sanford)
- A successful decade for Mendel’s Accountant
- Jacob’s livestock
- What would count as ‘new information’ in genetics?
- Fitness and ‘reductive evolution’
- Mutations, and why you shouldn‘t marry your cousin
- More evidence for the reality of genetic entropy
- Reply to Earl Rodd on on More evidence for the reality of genetic entropy
- Mutation accumulation rates are consistent with biblical creation
- Genetic entropy and simple organisms
- Genetic entropy and human lifespans
Human genetic history
- African origins and the rise of carnivory
- Patriarchal drive in the early post-Flood population
- Effective population sizes and loss of diversity during the Flood bottleneck
- Blue eyes mutation
- The genetic effects of the population bottleneck associated with the Genesis Flood
- Adam and Eve, designed diversity, and allele frequencies (8th International Conference on Creationism, 2018)
- An overview of the independent histories of the human Y chromosome and the human mitochondrial chromosome (8th International Conference on Creationism, 2018)
- Skin colour surprises
- Genetics questions answered
- Modelling biblical human population growth
- In light of genetics … Adam, Eve, and the creation/fall (Christian Apologetics Journal 12:51–98, 2014)
- Inbreeding and the origin of races
- Are women genetically superior to men?
- Is ‘mitochondrial Eve’ consistent with the biblical Eve?
- Could Adam and Eve have given rise to all the ‘races’?
- The Non-Mythical Adam and Eve! Refuting errors by Francis Collins and BioLogos
- Y-chromosome Adam and Cambrian explosion
- The chimpanzee Y chromosome is radically different from human
- A gentle answer; and the latest on “mitochondrial Eve”
- The Neutral Model of evolution and recent African origins
- The ‘Eve’ Mitochondrial Consensus Sequence (6th International Conference on Creationism, 2008)
- Species were designed to change, part 2
- Species were designed to change, part 1
- The human genome is amazingly complex: Massive new GTEx study counters Darwinism
- Genetic Diversity on Noah’s Ark
- More evidence for the reality of genetic entropy—update
- Natural Selection in Paradise
- Coronaviruses in creation
- Hachimoji DNA argues against evolution, despite recent claims
- Weird synthetic proteins are no help to evolution. Changing the genetic code points to design
- New sugar transport gene evolved in yeast?
- Barcodes, unclean animals, and skeletal mutations
- Can biologically active sequences come from random DNA?
- The four dimensional human genome defies naturalistic explanations
- Can mutations create new information?
- CMI scientific blunder?
- Splicing and dicing the human genome: Scientists begin to unravel the splicing code
- The slow, painful death of junk DNA
- Harnessing God’s design to help prevent sickness, but will the new vaccine technology alter our DNA?
- Unnatural selection: CRISPR on Netflix
- Gene editing babies? A dangerous, pointless experiment
- Human/animal hybrids?
- Human Cloning?
- Mammoth clones coming to a zoo near you
- The sophisticated Neandertal
- Are Neandertals pre-Flood people?
- Neandertal genome like ours (There may be Neandertals at your next family reunion!)
- The Painted Neandertal: Ancient cosmetics are upsetting evolutionary stories
- The Neandertal mitochondrial genome does not support evolution
- Taking a crack at the Neandertal mitochondrial genome
- Where did the Israelites cross the “Red Sea”?
- Göbekli Tepe shows evidence of geometric planning with Phil Robinson
- An Ancient textile factory?
- How does Göbekli Tepe fit with biblical history?
- Paleozoic Corals and Lunar Recession
- Coral: The animal that acts like a plant, but is an active predator, and makes its own rocks for a house
- ‘Ancient’ coral growth layers
- Reply to Daniel Jackson on ‘Ancient’ coral growth layers: yearly or monthly?
- Historical Science, Chaos Theory, and the sliding scale of trust with Paul Price
- Do animals have spirits? with Lita Sanders
- Answering question about 5G and COVID-19
- An open letter to Rhett McLaughlin: and anyone else on the road to unbelief
- Dystopian science: Part 1: Why the Bible enables science to work
- Dystopian science: Part 2: Conspiracy theories require a magical world
- Dystopian science: Part 3: Rebuilding science from the ground up
- We are less than dust
- How to think (not what to think)
- Why CMI rejects ‘conspiracy’ theory
- Emulating the mind of Christ in an age of misinformation
- Slaying yesterday’s dragons
- Five concise responses to atheistic arguments
- Time: The Great Enabler
- CMI ministers in the Caribbean
- Which of the physical sciences does CMI believe in?
- How did the earth dry out after the Flood?
- Answering coronavirus and flat earth questions
- Why the Universe does not revolve around the Earth: Refuting absolute geocentrism
- Refuting geocentrism response
- Refuting flat earth nonsense
- A direct test of the flat earth model: flight times
- Total eclipse of the brain
- Silicon based bugs: Scientists discover the first silicon-based life forms … in their imagination!
- The majestic gorilla with Lita Sanders
- The weird and wonderfully-designed sawfish
- Platypus thumbs its nose (or bill) at evolutionary scientists
- A long-overdue review of Hunter’s A Civic Biology
- Scopes at 100
- Yale university professor renounces Darwinism!
- Do these skulls prove common ancestry between apes and humans?
- Darwin’s Lamarckism vindicated?
- Darwin’s dying legacy
- NCSE Gives ‘Favorable’ Review of The Voyage the Shook the World
- Genetics and geographical distribution
- NASA scientist comes face to face with his Creator (Interview of rocket scientist Dr Henry Richter)
- From Atheism to Christ (Interview of molecular biologist Dr Yingguang Liu)
- Geneticist praises the Creator (Interview of ICR scientistDr Jeffrey Tomkins)
- Aerospace Engineer professes creation (Interview of Dr Dewey Hodges)
- Corals, genes and creation (Interview of Dr Carter)
- Evolutions’ Achilles’ Heels: A unique new book and DVD expose evolution’s fatal flaws (Interview of Dr Carter and Gary Bates)
- A long-overdue review of Hunter’s A Civic Biology (review of A Civic Biology George William Hunter
- Far-fetched speculations fail to replace the straight-forward understanding of Adam and Eve (review of The Genealogical Adam and Eve: The surprising science of universal ancestry, by S. Joshua Swamidass
- Reading evolution into the scriptures (review of Adam and the Genome: Reading scripture after genetic science, by Dennis R. Venema and Scot McKnight
- Finally setting the record straight—Christian views on the historicity of Adam through the ages (review of The Quest for the Historical Adam: Genesis, Hermeneutics, and Human Origins by William VanDoodewaard)
- A troubling thesis—Nicholas Wade pushes an old view of the origin of races (review of A Troublesome Inheritance: Genes, Race and Human History, by Nicholas Wade).
July 3, 2021

Matthew Cserhati, Ph.D., Bioinformatics, gave a LIVE, IN-PERSON presentation titled, "MOLECULAR BARAMINOLOGY." "Baraminology" refers to the science of organizing today's plants and animals according to their inferred created "kinds," ("baramin" in Hebrew). "Molecular baraminology" seeks to resolve each Genesis kind by identifying multiple distinctive molecules shared by members of the kind. Dr. Cserhati has developed algorithms that allow him to rapidly sift through huge genetic data banks, identifying created kinds by shared nuclear DNA, shared mitochondrial DNA, and and shared gene sequences.
To see his presentation, click HERE.
About our speaker (The following information is borrowed from Creation Ministries International's website at https://creation.com/dr-matthew-cserhati-cv.)
Education
During his Ph.D. studies, Matthew developed a transcription factor binding site dyad prediction algorithm, with application to crop species at the Biological Research Center in Szeged, Hungary. He also developed a dedicated website for housing haplotype information on candidate drought genes in barley and taught two science courses.
From 2010–2011, he took part in the development of the program OPTIMA in the laboratory of Chemical Kinetics in the Institute of Chemistry at ELTE University, in Budapest. This work involved the estimation of a number of Arrhenius parameters in a number of combustion reactions. He also taught Linux programming as a special course.
From 2011–2013, he worked as a post-doctoral research fellow at the University of Nebraska, Lincoln. The two main projects he was involved with included the analysis of Next-gen sequencing data to discover differentially expressed genes in the green algae, Chlamydomonas reinhardtii due to low carbon concentrations, and the analysis of the regulation of 177 ribosomal protein genes across ~30,000 experiments, representing hundreds of gigabytes of data from the ENCODE project.
From 2013–2017, he worked as a bioinformatics programmer at the University of Nebraska Medical Center. His tasks included the development of the NNTC-DCC database, a browsable, integrated website for storing experimental and clinical data on neuroAIDS patients. He was also involved in the demultiplexing, storage, and analysis of Next-gen sequencing data for wet-lab researchers. Other notable projects included the de novo genome assembly and annotation of ten NCLDV (NucleoCytoplasmic Large DNA Viruses), as well as the application of a motif prediction algorithm to the genomes of modern and archaic humans. He also helped teach a couple courses in bioinformatics.
In 2017 he started an M.A. program at Greenville Presbyterian Theological Seminary.
From 2018–2019, he took part in the prediction and analysis of differentially expressed genes in hepatocellular carcinoma (HCC) in mouse and human at the University of Texas Health Science Center at San Antonio.
From 2010 He performed over 200 translations and interpretation jobs in Hungarian, German, Swedish, Dutch and Norwegian. He was also a member of the American Translators Association and the Nebraska Association for Translators and Interpreters for several years and he led an informal biblical languages club at Grace Orthodox Presbyterian Church in San Antonio, Texas.
In 2001 he helped establish a creation science group in Hungary called the Protestant Creation Research Group. In 2015 he became a member of the Creation Research Society.
Creationist publications and articles (using several pseudonyms)
To see his presentation, click HERE.
About our speaker (The following information is borrowed from Creation Ministries International's website at https://creation.com/dr-matthew-cserhati-cv.)
Education
- 2011, University of Szeged, Hungary, Ph.D., Bioinformatics
- 2010, University of Szeged, Hungary, B.Sc., Software Development
- 2003, Eötvös Loránd University, M.Sc., Biology
- Native Hungarian speaker
- Swedish, Advanced level, complex, oral
- Dutch, Intermediate level, oral + written
- German, Intermediate level, oral
- Norwegian, Intermediate level, written
- Beginning Greek, Greenville Presbyterian Theological Seminary
- Hebrew I+II, Greenville Presbyterian Theological Seminary
During his Ph.D. studies, Matthew developed a transcription factor binding site dyad prediction algorithm, with application to crop species at the Biological Research Center in Szeged, Hungary. He also developed a dedicated website for housing haplotype information on candidate drought genes in barley and taught two science courses.
From 2010–2011, he took part in the development of the program OPTIMA in the laboratory of Chemical Kinetics in the Institute of Chemistry at ELTE University, in Budapest. This work involved the estimation of a number of Arrhenius parameters in a number of combustion reactions. He also taught Linux programming as a special course.
From 2011–2013, he worked as a post-doctoral research fellow at the University of Nebraska, Lincoln. The two main projects he was involved with included the analysis of Next-gen sequencing data to discover differentially expressed genes in the green algae, Chlamydomonas reinhardtii due to low carbon concentrations, and the analysis of the regulation of 177 ribosomal protein genes across ~30,000 experiments, representing hundreds of gigabytes of data from the ENCODE project.
From 2013–2017, he worked as a bioinformatics programmer at the University of Nebraska Medical Center. His tasks included the development of the NNTC-DCC database, a browsable, integrated website for storing experimental and clinical data on neuroAIDS patients. He was also involved in the demultiplexing, storage, and analysis of Next-gen sequencing data for wet-lab researchers. Other notable projects included the de novo genome assembly and annotation of ten NCLDV (NucleoCytoplasmic Large DNA Viruses), as well as the application of a motif prediction algorithm to the genomes of modern and archaic humans. He also helped teach a couple courses in bioinformatics.
In 2017 he started an M.A. program at Greenville Presbyterian Theological Seminary.
From 2018–2019, he took part in the prediction and analysis of differentially expressed genes in hepatocellular carcinoma (HCC) in mouse and human at the University of Texas Health Science Center at San Antonio.
From 2010 He performed over 200 translations and interpretation jobs in Hungarian, German, Swedish, Dutch and Norwegian. He was also a member of the American Translators Association and the Nebraska Association for Translators and Interpreters for several years and he led an informal biblical languages club at Grace Orthodox Presbyterian Church in San Antonio, Texas.
In 2001 he helped establish a creation science group in Hungary called the Protestant Creation Research Group. In 2015 he became a member of the Creation Research Society.
Creationist publications and articles (using several pseudonyms)
- Cserhati, M., Baraminology data filtering method based on entropy measurement and its application in dinosaur and cephalopod data sets, Journal of Creation 33(3):55-65, 2019.
- Cserhati, M., Alquist, J., Baraminology suggests cryptic relationships among Caprimulgiformes, Journal of Creation 33(3):66-76, 2019.
- Cserhati, M., and Tay, J., Comparison of morphology-based and genomics-based baraminology methods. Journal of Creation 33(3):49-55, 2019.
- Cserhati, M., The green algae Chlamydomonas reinhardtii finds safety in numbers by design, J Creation 33(2):93-98, 2019.
- Cserhati, M., and Chira, A., Critique of the latest evolutionary models of the beginning of life. Journal of Creation 33(1):78–84, 2019.
- O’Micks, J., Baraminic analysis of archaic and modern human genomes. Journal of Creation 32(3):19–24, 2018.
- Cserhati, M., Swinging too far to the other side. Journal of Creation 32(2):60–62, 2018.
- Cserhati, M., What does the catechism of the Roman Catholic Church say about creation? Journal of Creation 32(2):79–82, 2018.
- Cserhati, M., and Shipton, W., The irreducibly complex ribosome is a unique creation in the three domains of life. Journal of Creation 31(2):94-102, 2017.
- O’Micks, J., Baraminic analysis of nucleocytoplasmic large DNA viruses. Journal of Creation 31(1):73-79, 2017.
- O’Micks, J., Creation perspective of nucleocytoplasmic large DNA viruses. Journal of Creation 30(3):110-117, 2016.
- O’Micks, J., Promoter evolution is impossible by random mutations. Journal of Creation 30(2):60-66, 2016.
- O’Micks, J., Pseudogenes and bacterial genome decay. Journal of Creation 30(1):104-110, 2016.
- O’Micks, J., Cnidarians turn evolutionary theory into jelly. Journal of Creation 29(3):71-79, 2015. See also the response in Journal of Creation 30(2):51, 2016.
- O’Micks, J., Bacterial genome decay from a baraminological viewpoint. Journal of Creation 29(2):122-130, 2015.
- Savanne, D., Denisovans menace evolution—a new chapter in the human origins debate. Journal of Creation 28(3):5-8, 2014.
- O’Micks, J., Small genome size of Utricularia gibba problematic for evolution but not creation. Journal of Creation 27(3):4-6, 2013.
- Arneigh, M.R., It’s a small world—microRNA cuts evolution down to size. Journal of Creation 27(2):85-90, 2013.
- Shan, L.E., Transposon amplification in rapid intrabaraminic diversification. Journal of Creation 23(2):110-117, 2009.
- Cserháti, M., Creation aspects of conserved non-coding sequences. Journal of Creation 21(2):101-108, 2007.
- O’Micks, J., A preliminary cephalopod baraminology study based on the analysis of mitochondrial genomes and morphological characteristics. Answers Research Journal 11:193–204, 2018.
- O’Micks, J., Further evidence that Homo naledi is not a member of the human holobaramin based on measurements of vertebrae and ribs. Answers Research Journal 10:103–113, 2017.
- O’Micks, J., Likely Discontinuity Between Humans and Non-Human Hominins Based on Endocranial Volume and Body Mass With a Special Focus on Homo naledi—A Short Analysis. Answers Research Journal 10:241-243, 2017.
- O’Micks, J., Rebuttal to “Reply to O’Micks concerning the geology and taphonomy of the Homo naledi site” and “Identifying humans in the fossil record: a further response to O’Micks”. Answers Research Journal 10:63–70, 2017.
- O’Micks, J., Reply to “Taxon sample in hominin baraminology: a response to O’Micks”. Answers Research Journal 9:373–375, 2016.
- O’Micks, J., Molecular structures shared by prokaryotes and eukaryotes show signs of only analogy and not homology. Answers Research Journal 9:284–292, 2016.
- O’Micks, J., Homo naledi probably not part of the human holobaramin based on baraminic re-analysis including postcranial evidence. Answers Research Journal 9:263–272, 2016.
- O’Micks, J., Baraminology classification based on gene content similarity measurement. Creation Research Society Quarterly 54:27–37, 2017.
- Yaugh, A., Baraminological analysis of a set of Archaea species based on genomic data. Creation Research Society Quarterly 53:140–154, 2016.
- O’Micks, J., Preliminary baraminological analysis of Homo naledi and its place within the human baramin. Journal of Creation Theology and Science Series B: Life Sciences 6: 31–39, 2016.
- More of Dawkins’ same old tired rhetoric. Review of Outgrowing God by Richard Dawkins
- O’Micks, J., Taking the bite out of evolution — critical review of Evolution’s Bite by Peter S. Ungar. Journal of Creation 31(3):24-27, 2017.
- O’Micks, J., Modern science catches up with Neandertal man, a review of The Neanderthals Rediscovered—How modern science is rewriting their story by Dimitra Papagianni and Michael A. Morse. Journal of Creation 32(1):38–42, 2017.
- O’Micks, J., and Clarey, T., Book Review of Almost Human, the astonishing tale of Homo naledi and the discovery that changed our human story, by Lee Berger and John Hawks. Answers Research Journal 10:187–194, 2017.
- Cserhati, M., Xiao, P., Guda, C., K-mer-Based Motif Analysis in Insect Species across Anopheles, Drosophila, and Glossina Genera and Its Application to Species Classification, Computational and Mathematical Methods in Medicine vol. 2019, 2019.
- Cserhati, M., Mooter, M.E., Fan, M., Wicks, B., Xiao, P., Pauley, M., and Guda, C., Motif comparison between human, Neandertals and Denisova. BMC Genomics 19(1):472, 2018 | doi:10.1186/s12864-018-4710-1.
- Lopez, W., Page, A.M., Carlson, D.J., Ericson, B.L., Cserhati, M.F., Guda, C., and Carlson, K.A., Analysis of immune-related genes during Nora virus infection of Drosophila melanogaster using next generation sequencing. AIMS Microbiology 4(1):123–139, 2018. doi: 10.3934/microbiol.2018.1.123.
- Cserhati, M., Motif content comparison between monocot and dicot species. Genom Data 3:128–36, 2015 | doi:10.1016/j.gdata.2014.12.006.
- Pettkó-Szandtner, A., Cserháti, M., Barrôco, R.M., Hariharan, S., Dudits, D., and Beemster, G.T., Core cell cycle regulatory genes in rice and their expression profiles across the growth zone of the leaf. J Plant Res 128(6):953—74, 2015 | doi:10.1007/s10265-015-0754-3.
- Cserhati, M., Pandey, S., Beaudoin, J.J., Baccaglini, L., Guda, C., and Fox, H.S., The National NeuroAIDS Tissue Consortium (NNTC) Database: an integrated database for HIV-related studies. Database (Oxford) 2015:bav074 | doi:10.1093/database/bav074.
- Szécsényi, M., Cserháti, M., Zvara, Á., Dudits, D., and Györgyey, J., Monitoring of transcriptional responses in roots of six wheat cultivars during mild drought stress. Cereal Research Communications 41(4):528–538, 2013 | doi:10.1556/CRC.41.2013.4.3.
- Cserháti, M., Turóczy, Z., Dudits, D., and Györgyey, J., The rice word landscape: a detailed catalogue of the rice motif content in the non-coding regions. OMICS 16(6):334–42, 2012. doi:10.1089/omi.2011.0056.
- Cserhati, M., Prommute – A promoter mutation simulation for modeling the evolution of genetic regulatory elements. Journal of Computer Science and Systems Biology 5:74–80, 2012 |doi:10.4172/jcsb.1000093.
- Brueggeman, A.J., Gangadharaiah, D.S., Cserhati, M., Diaz-Cano, D.C., Weeks, D.P., and Ladunga, I., Activation of the carbon concentrating mechanism by CO2 deprivation coincides with massive transcriptional restructuring in Chlamydomonas reinhardtii. Plant Cell 24(5):1860–75, 2012 | doi:10.1105/tpc.111.093435.
- Zsély, I.,Gy., Varga, T., Nagy, T., Cserháti, M., Turányi, T., Peukert, S., Braun-Unkhoff, M., Naumann, C., and Riedel, U., Determination of rate parameters of cyclohexane and 1-hexene decomposition reactions. Energy 43(1):85–93, 2012 | doi:10.1016/j.energy.2012.01.004.
- Turányi, T., Nagy, T., Zsély, I.Gy., Cserháti, M., Varga, T., Szabó, B.T., Sedyó, I., Kiss, P.T., Zempléni, A., and Curran, H.J, Determination of rate parameters based on both direct and indirect measurements. Int J Chem Kinet 44(5):284–302, 2012 | doi:10.1002/kin.20717.
- Áy, Z., Mihály, R., Cserháti, M., Kótai, É., and Pauk, J., The effect of high concentrations of glufosinate ammonium on the yield components of transgenic spring wheat (Triticum aestivum L.) constitutively expressing the bar gene. The Scientific World Journal 2012:657945 | doi:10.1100/2012/657945.
- Cseri, A., Cserháti, M., von Korff, M., Bettina, N., Gábor, V.H., Palágyi, A., Pauk, J., Dudits, D., and Törjék, O., Allele mining and haplotype discovery in barley candidate genes for drought tolerance. Euphytica 181:341, 2012 | doi:10.1007/s10681-011-0445-7.
- Cserháti, M., Turóczy, Z., Zombori, Z., Cserzo, M., Dudits, D., Pongor, S., and Györgyey, J., Prediction of new abiotic stress genes in Arabidopsis thaliana and Oryza sativa according to enumeration-based statistical analysis. Mol Genet Genomics 285(5):375–91, 2011 | doi:10.1007/s00438-011-0605-4.
- Turóczy, Z., Kis, P., Török, K., Cserháti, M., Lendvai, A., Dudits, D., and Horváth, G.V., Overproduction of a rice aldo-keto reductase increases oxidative and heat stress tolerance by malondialdehyde and methylglyoxal detoxification. Plant Mol Biol 75(4–5):399–412, 2011 | doi:10.1007/s11103-011-9735-7.
- Cserhati, M., and Gyorgyey, J., “Gene research in silico”, in The science of wheat breeding (ed. Denes Dudits), Winter Fair Ltd., Eger, 2006.
- Dudits, D., Cserháti, M., Miskolczi, P., and Horváth, G., “The growing family of plant cyclin dependent kinases with multiple functions in cellular and developmental regulation”, in Cell cycle control and development (ed. Dirk Inzé), Blackwell Publishing, Oxford, 2006.
- Baraminology suggests cryptic relationships among Caprimulgiformes with Jon Ahlquist. From Journal of Creation 33(3), 2019.
- Baraminology data filtering method based on entropy measurement and its application in dinosaur and cephalopod data sets from Journal of Creation 33(3), 2019.
- Comparison of morphology-based and genomics-based baraminology methods with Joel Tay. From Journal of Creation 33(3), 2019.
- One for all and all for one from Creation magazine 41(1) 2020.
- The wonderful world of bats from Creation magazine 41(1) 2020.
- The wonderfully designed cell cycle from Creation magazine 41(4) 2019.
- Evidence that modern humans and ‘super-archaic’ humans may have shared DNA
- What should Christians think about artificial selection and genetic modification?
- Jacob’s livestock
- More of the moth myth!
- Owls—masters of the night sky from Creation magazine 42(3) 2020.
- The priests of Darwin want your children
- Cosmic Biology with Paul Price
- Is the RubisCO enzyme an ineffective leftover of evolution?
- Snakes losing legs is not evolution!
- Do koalas prove that humans got part of their DNA from viruses?
- Racial mixing is perfectly biblical!
- Is a brain a human being?
- Weird synthetic proteins are no help to evolution. Changing the genetic code points to design
- What is Homo luzonensis?
- Evolution of middle ear bones in mammals from jaw bones in reptiles?
- Yale university professor renounces Darwinism!
- Reformation wall vandalized yet again
- From a rib to a 206-bone human
- Can mutations lead to new genetic information?
- Is the Shroud of Turin authentic?
- Researching creation after secular academia
- Rails derail evolution
- Ambopteryx – bird or bat-winged reptile?
- New sugar transport gene evolved in yeast?
- Were stem cell-like organisms the first forms of life?
- Did the ear bones of mammals really evolve from the jawbones of reptiles?
- Homeschool conference: great encouragement and some concerns
- Hachimoji DNA argues against evolution, despite recent claims
- Newly discovered Medusavirus turns evolutionary theory to stone
- Have astrobiologists really found a super-Earth?
- What is Archaeotherium? Pig or hippopotamus? What is Onychonycteris? Shrew or bat?
- More of Dawkins’ same old tired rhetoric - Review of Outgrowing God by Richard Dawkins
June 19, 2021

We watched the video,
WORLD RENOWN DOCTOR BLOWS THE LID OFF OF COVID VACCINE
https://rumble.com/vhp7y5-full-interview-world-renowned-doctor-blows-lid-off-of-covid-vaccine.html
About the Video's Featured Speaker:
Dr. Peter McCullough has been the world's most prominent and vocal advocate for early outpatient treatment of SARS-CoV-2 (COVID-19) Infection in order to prevent hospitalization and death. On May 19, 2021, he was interviewed regarding his efforts as a treating physician and researcher. From his unique vantage point, he has observed and documented a PROFOUNDLY DISTURBING POLICY RESPONSE to the pandemic -- a policy response that may prove to be the greatest malpractice and malfeasance in the history of medicine and public health.
Dr. McCullough is an internist, cardiologist, epidemiologist, and Professor of Medicine at Texas A & M College of Medicine, Dallas, TX USA. Since the outset of the pandemic, Dr. McCullough has been a leader in the medical response to the COVID-19 disaster and has published “Pathophysiological Basis and Rationale for Early Outpatient Treatment of SARS-CoV-2 (COVID-19) Infection” the first synthesis of sequenced multidrug treatment of ambulatory patients infected with SARS-CoV-2 in the American Journal of Medicine and subsequently updated in Reviews in Cardiovascular Medicine. He has 40 peer-reviewed publications on the infection and has commented extensively on the medical response to the COVID-19 crisis in TheHill and on FOX NEWS Channel. On November 19, 2020, Dr. McCullough testified in the US Senate Committee on Homeland Security and Governmental Affairs and throughout 2021 in the Texas Senate Committee on Health and Human Services, Colorado General Assembly, and New Hampshire Senate concerning many aspects of the pandemic response.
Peter A. McCullough, MD, MPH, FACP, FACC, FAHA, FCRSA, FCCP, FNKF, FNLA
Professor of Medicine, Texas A & M College of Medicine
Board Certified Internist and Cardiologist
President Cardiorenal Society of America
Editor-in-Chief, Reviews in Cardiovascular Medicine
Editor-in-Chief, Cardiorenal Medicine
Senior Associate Editor, American Journal of Cardiology
For more information about Dr. McCullough, please visit: heartplace.com/dr-peter-a-mccullough
June 5, 2021

Dennis Petersen, M.S., gave a presentation titled, TRANSMUTATIONS? DO SIMPLE FORMS REALLY EVOLVE INTO COMPLEX FORMS OF LIFE OVER TIME?
About Our Speaker:
Dennis R. Petersen is the author of the Unlocking the Mysteries of Creation seminar, book, and video curriculum. Dennis founded and has served as the President of the Creation Resource Foundation since 1983. He is also the host of the Sacramento Radio Show, Reclaiming Your Legacy, on KFIA am710. He has conducted creation seminars in churches throughout America and in other countries for the past three decades.
The Creation Resource Foundation, begun in 1983 by former museum curator and pastor, Dennis Petersen, is the providential result of a vision Dennis shares to communicate the gospel of Jesus Christ to our information-flooded, media-confused generation.
To view the recorded presentation, click here: https://drive.google.com/drive/folders/11ky8DYyFEwCLrK7MA_III_l2z6Ucnti1
About Our Speaker:
Dennis R. Petersen is the author of the Unlocking the Mysteries of Creation seminar, book, and video curriculum. Dennis founded and has served as the President of the Creation Resource Foundation since 1983. He is also the host of the Sacramento Radio Show, Reclaiming Your Legacy, on KFIA am710. He has conducted creation seminars in churches throughout America and in other countries for the past three decades.
The Creation Resource Foundation, begun in 1983 by former museum curator and pastor, Dennis Petersen, is the providential result of a vision Dennis shares to communicate the gospel of Jesus Christ to our information-flooded, media-confused generation.
To view the recorded presentation, click here: https://drive.google.com/drive/folders/11ky8DYyFEwCLrK7MA_III_l2z6Ucnti1
MAY 15, 2021

We watched several short videos demonstrating the presence of a larger abstract universe, one which overarches and influences the material cosmos of matter and energy. These included FIBONACCI AND THE GOLDEN MEAN ( 27 min, 13 sec);
We watched several short videos demonstrating the presence of a larger abstract universe, one which overarches and influences the material cosmos of matter and energy. These included FIBONACCI AND THE GOLDEN MEAN ( 27 min, 13 sec);



FIBONACCI NUMBERS PUT TO MUSIC (2 min, 40 sec)
FIBONACCI NUMBERS PUT TO MUSIC (2 min, 40 sec)

Finally, we watched PRIVILEGED SPECIES, ( 32 min,
15 sec), which discusses the fine-tuning of the universe to make life for humankind possible.
Finally, we watched PRIVILEGED SPECIES, ( 32 min,
15 sec), which discusses the fine-tuning of the universe to make life for humankind possible.
May 1, 2021

We watched a video by Dr. Jason Lisle titled, "THE SECRET CODE OF CREATION."
Showcasing Mandelbrot fractals as evidence, Dr. Lisle demonstrated that there is more to reality than the material universe of matter and energy. Governing the physical cosmos lies a universe of abstract entities, patterns, and relationships that is unarguably REAL, that secular science cannot explain. This abstract universe reflects the unfathomable Genius of our Creator.
"THE SECRET CODE OF CREATION" is currently available on YouTube.
Showcasing Mandelbrot fractals as evidence, Dr. Lisle demonstrated that there is more to reality than the material universe of matter and energy. Governing the physical cosmos lies a universe of abstract entities, patterns, and relationships that is unarguably REAL, that secular science cannot explain. This abstract universe reflects the unfathomable Genius of our Creator.
"THE SECRET CODE OF CREATION" is currently available on YouTube.
Proudly powered by Weebly